Residues substitution in the active site of DSBA may compensate for the lack of the canonical motif CPHC
Rinaldi, F.C., Guimaraes, B.G.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Disulfide oxidoreductase | 193 | Xylella fastidiosa | Mutation(s): 0 Gene Names: XF1436 EC: 1.8.4.2 | 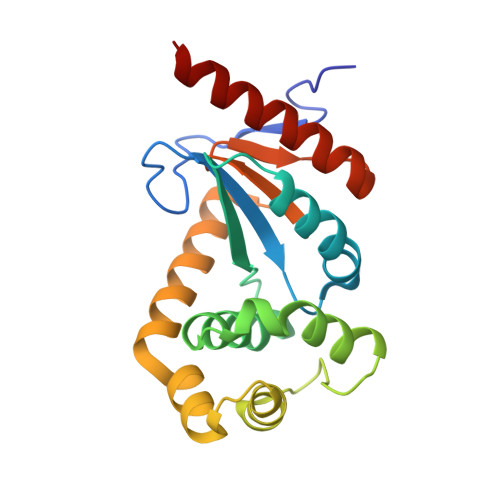 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q9PDE3 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| 8 residue peptide | D [auth T] | 8 | Xylella fastidiosa | Mutation(s): 0 | 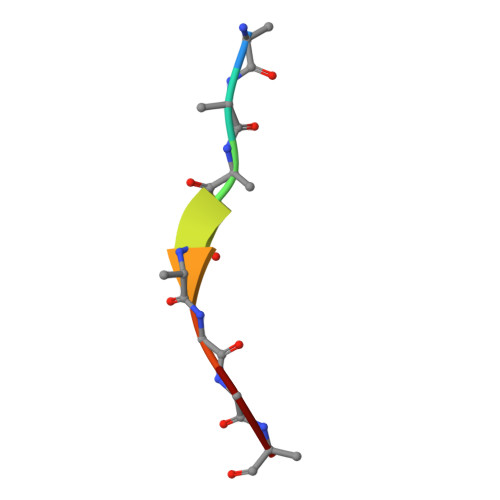 |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 200.117 | α = 90 |
| b = 41.722 | β = 95.87 |
| c = 79.807 | γ = 90 |
| Software Name | Purpose |
|---|---|
| DENZO | data reduction |
| SCALEPACK | data scaling |
| SHARP | phasing |
| REFMAC | refinement |
| PDB_EXTRACT | data extraction |
| CBASS | data collection |
| HKL-2000 | data reduction |
| HKL-2000 | data scaling |