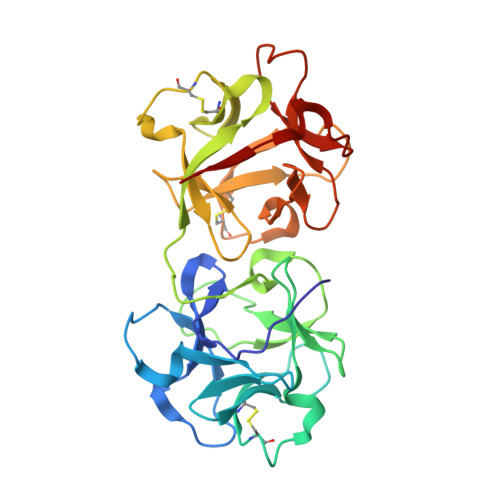
The mistletoe lectin I--phloretamide structure reveals a new function of plant lectins.
Meyer, A., Rypniewski, W., Celewicz, L., Erdmann, V.A., Voelter, W., Singh, T.P., Genov, N., Barciszewski, J., Betzel, C.h.(2007) Biochem Biophys Res Commun 364: 195-200
- PubMed: 17937929 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2007.09.113
- Primary Citation Related Structures:
2R9K - PubMed Abstract:
The X-ray structure at 2.7A resolution of the complex between the European mistletoe lectin I (Viscum album, ML-I) and the plant growth hormone, 3-(p-hydroxyphenyl)-propionic acid amide (phloretamide, PA) from xylem sap has revealed the binding of PA at the so far undescribed hydrophobic cavity located between the two subunits of this ribosome-inhibiting protein. No such cavity is observed in related lectins. The binding of PA is achieved through interactions with the non-conserved residues Val228A, Leu230A, Arg388B, and the C-terminal Pro510B. It is conceivable that binding of PA to ML-I is part of a defence mechanism of the parasite against the host, whereby the parasite prevents the growth hormone of the host from interfering with its own regulatory system. The specific binding of PA to ML-I indicates that heterodimeric RIPs are multifunctional proteins whose functions in the cell have not yet been fully recognized and analyzed.
- Institute of Biochemistry and Molecular Biology, University of Hamburg, c/o DESY, Notkestr. 85, Building 22a, 22603 Hamburg, Germany.
Organizational Affiliation: