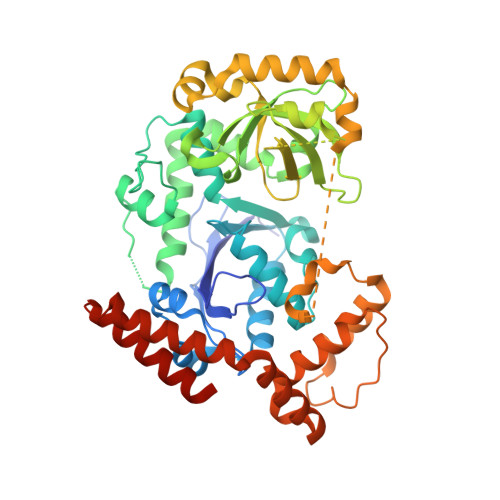
Structure of Escherichia coli exonuclease I in complex with thymidine 5'-monophosphate.
Busam, R.D.(2008) Acta Crystallogr D Biol Crystallogr 64: 206-210
- PubMed: 18219121 Search on PubMed
- DOI: https://doi.org/10.1107/S090744490706012X
- Primary Citation Related Structures:
2QXF - PubMed Abstract:
In Escherichia coli, exonuclease I (ExoI) is a monomeric processive 3'-5' exonuclease that degrades single-stranded DNA. The enzyme has been implicated as primarily being involved in repairing frameshift mutations. The structure of the enzyme has previously been determined in a hexagonal space group at 2.4 A resolution. Here, the structure of ExoI in complex with a nucleotide product, thymidine 5'-monophosphate, is described in an orthorhombic space group at 1.5 A resolution. This new high-resolution structure provides some insight into the interactions involved in binding a nucleotide product. The conserved active site contains a unique metal-binding position when compared with orthologous sites in the Klenow fragment, T4 DNA polymerase and dnaQ. This unique difference is proposed to be a consequence of the repositioning of an important histidine, His181, away from the active site.
- Institute of Molecular Biology, Howard Hughes Medical Institute and Department of Physics, 1229 University of Oregon, Eugene, OR 97403-1229, USA. robert.busam@ki.se
Organizational Affiliation: