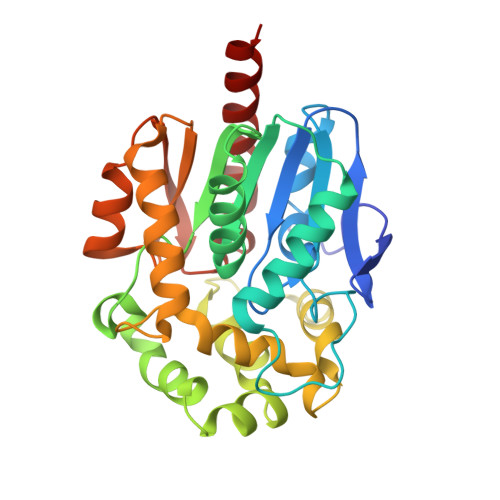
X-ray crystal structure of Mycobacterium tuberculosis haloalkane dehalogenase Rv2579.
Mazumdar, P.A., Hulecki, J.C., Cherney, M.M., Garen, C.R., James, M.N.(2008) Biochim Biophys Acta 1784: 351-362
- PubMed: 18062934 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbapap.2007.10.014
- Primary Citation Related Structures:
2QVB - PubMed Abstract:
Haloalkane dehalogenases are enzymes well known to be important in bioremediation; the organisms from which they are produced are able to clean up toxic organohalides from polluted environments. However, besides being found in such contaminated environments, these enzymes have also been found in root or tissue-colonizing bacterial species. The haloalkane dehalogenase Rv2579 from Mycobacterium tuberculosis H37Rv has been cloned, expressed, purified and its crystal structure determined at high resolution (1.2A). In addition, the crystal structure of the enzyme has been determined in complex with the product from the reaction with 1,3-dibromopropane, i.e. 1,3-propanediol and in complex with the classical substrate of haloalkane dehalogenases, 1,2-dichloroethane. The enzyme is a two-domain protein having a catalytic domain of an alpha/beta hydrolase fold and a cap domain. The active site residues and the halide-stabilizing residues have been identified as Asp109, Glu133, His273, Asn39 and Trp110. Its overall structure is similar to those of other known haloalkane dehalogenases. Its mechanism of action involves an SN2 nucleophilic displacement.
- Group in Protein Structure and Function, Department of Biochemistry, University of Alberta, Edmonton, Alberta, Canada T6G 2H7.
Organizational Affiliation: