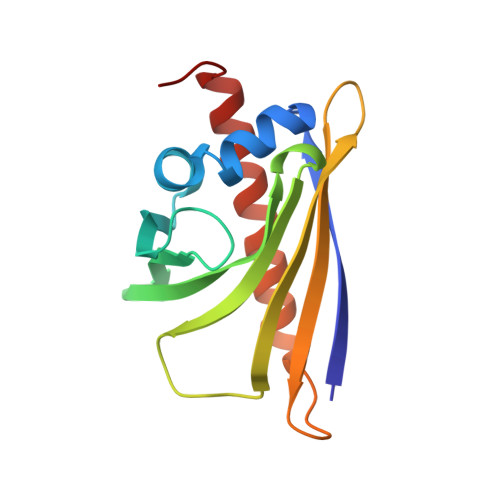
Lupinus luteus pathogenesis-related protein as a reservoir for cytokinin.
Fernandes, H., Pasternak, O., Bujacz, G., Bujacz, A., Sikorski, M.M., Jaskolski, M.(2008) J Mol Biology 378: 1040-1051
- PubMed: 18406424 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2008.03.027
- Primary Citation Related Structures:
2QIM - PubMed Abstract:
Plant pathogenesis-related (PR) proteins of class 10 (PR-10) are small and cytosolic. The main feature of their three-dimensional structure is a large cavity between a seven-stranded antiparallel beta-sheet and a long C-terminal alpha-helix. Although PR-10 proteins are abundant in plants, their physiological role remains unknown. Recent data have indicated ligand binding as their possible biological function. The article describes the structure of a complex between a classic PR-10 protein (yellow lupine LlPR-10.2B) and the plant hormone, trans-zeatin. Previously, trans-zeatin binding has been reported in a structurally related cytokinin-specific binding protein, which has a distant sequence relation with classic PR-10 proteins. In the present 1.35 A resolution crystallographic model, three perfectly ordered zeatin molecules are found in the binding cavity of the protein. The fact that three zeatin molecules are bound by the protein when only a fourfold molar excess of the ligand was used indicates an unusual type of affinity for this ligand and suggests that LlPR-10.2B, and perhaps other PR-10 proteins as well, acts as a reservoir of cytokinin molecules in the aqueous environment of the cell.
- Center for Biocrystallographic Research, Institute of Bioorganic Chemistry, Polish Academy of Sciences, 61-704 Poznan, Poland.
Organizational Affiliation: