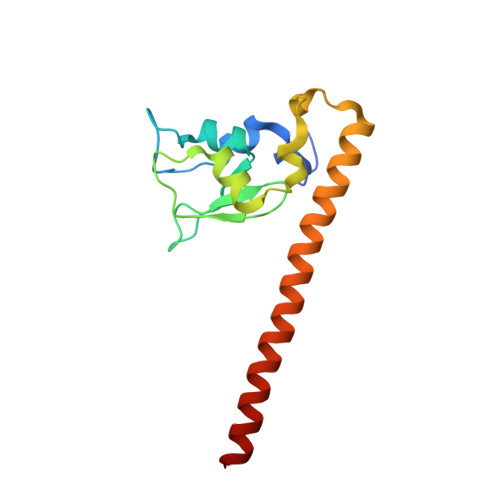
Structure of a Survivin-Borealin-INCENP Core Complex Reveals How Chromosomal Passengers Travel Together.
Jeyaprakash, A.A., Klein, U.R., Lindner, D., Ebert, J., Nigg, E.A., Conti, E.(2007) Cell 131: 271-285
- PubMed: 17956729 Search on PubMed
- DOI: https://doi.org/10.1016/j.cell.2007.07.045
- Primary Citation Related Structures:
2QFA - PubMed Abstract:
The chromosomal passenger complex (CPC) is a key regulator of chromosome segregation and cytokinesis. CPC functions are connected to its localization. The complex first localizes to centromeres and later associates with the central spindle and midbody. Survivin, Borealin, and INCENP are the three components of the CPC that regulate the activity and localization of its enzymatic component, the kinase Aurora B. We determined the 1.4 A resolution crystal structure of the regulatory core of the CPC, revealing that Borealin and INCENP associate with the helical domain of Survivin to form a tight three-helical bundle. We used siRNA rescue experiments with structure-based mutants to explore the requirements for CPC localization. We show that the intertwined structural interactions of the core components lead to functional interdependence. Association of the core "passenger" proteins creates a single structural unit, whose composite molecular surface presents conserved residues essential for central spindle and midbody localization.
- Max-Planck-Institute of Biochemistry, Am Klopferspitz 18, D-82152 Martinsried, Germany.
Organizational Affiliation: