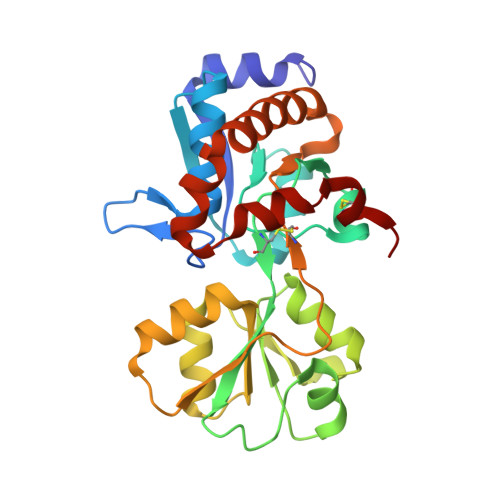
Crystal structure of the ligand-binding protein EhuB from Sinorhizobium meliloti reveals substrate recognition of the compatible solutes ectoine and hydroxyectoine.
Hanekop, N., Hoing, M., Sohn-Bosser, L., Jebbar, M., Schmitt, L., Bremer, E.(2007) J Mol Biology 374: 1237-1250
- PubMed: 17996893 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2007.09.071
- Primary Citation Related Structures:
2Q88, 2Q89 - PubMed Abstract:
In microorganisms, members of the binding-protein-dependent ATP-binding cassette transporter superfamily constitute an important class of transport systems. Some of them are involved in osmoprotection under hyperosmotic stress by facilitating the uptake of "compatible solutes". Currently, the molecular mechanisms used by these transport systems to recognize compatible solutes are limited to transporters specific for glycine betaine and proline betaine. Therefore, this study reports a detailed analysis of the molecular principles governing substrate recognition in the Ehu system from Sinorhizobium meliloti, which is responsible for the uptake of the compatible solutes ectoine and hydroxyectoine. To contribute to a broader understanding of the molecular interactions underlying substrate specificity, our study focused on the substrate-binding protein EhuB because this protein binds the ligand selectively, delivers it to the translocation machinery in the membrane and is thought to be responsible for substrate specificity. The crystal structures of EhuB, in complex with ectoine and hydroxyectoine, were determined at a resolution of 1.9 A and 2.3 A, respectively, and allowed us to assign the structural principles of substrate recognition and binding. Based on these results, site-directed mutagenesis of amino acids involved in ligand binding was employed to address their individual contribution to complex stability. A comparison with the crystal structures of other binding proteins specific for compatible solutes revealed common principles of substrate recognition, but also important differences that might be an adaptation to the nature of the ligand and to the demands on protein affinity imposed by the environment.
- Institute of Biochemistry, Heinrich Heine University Duesseldorf, Universitaetsstrasse 1, 40225 Duesseldorf, Germany.
Organizational Affiliation: