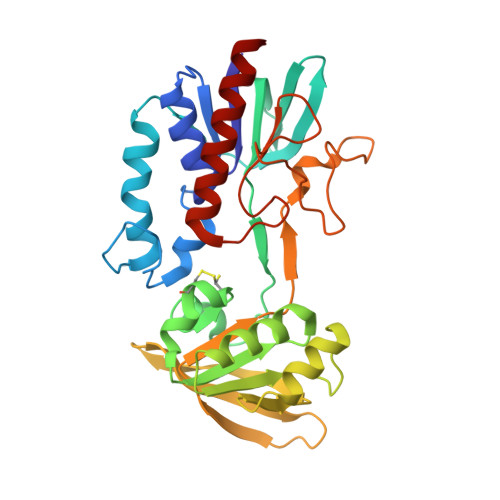
High-resolution structures of oxidized and reduced thioredoxin reductase from Helicobacter pylori.
Gustafsson, T.N., Sandalova, T., Lu, J., Holmgren, A., Schneider, G.(2007) Acta Crystallogr D Biol Crystallogr 63: 833-843
- PubMed: 17582174 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444907026303
- Primary Citation Related Structures:
2Q0K, 2Q0L - PubMed Abstract:
The crystal structures of homodimeric thioredoxin reductase (TrxR) from the gastric pathogen Helicobacter pylori in complex with NADP(+) have been determined for the oxidized and reduced form of the enzyme at 1.7 and 1.45 A resolution, respectively. The enzyme subunit is built up of FAD- and NAD(P)H-binding domains, each of which contain variants of the Rossmann fold typical of nucleotide-binding proteins. The majority of the amino-acid residues binding the cofactors FAD and NAD(P)H are conserved in the low-molecular-weight thioredoxin reductases. In the reduced species the isoalloxazine ring of FAD is bent along an axis passing through the N5 and N10 atoms with an angle of 22 degrees and the ribityl moiety adopts an unusual conformation. Well defined electron density shows the position of the whole NADP(+) molecule with a binding mode similar to that observed in the Escherichia coli TrxR-thioredoxin complex, although the orientation of the NAD(P)H-binding domain is different. Rigid-body rotation of this domain to the orientation observed in the E. coli TrxR-thioredoxin complex positions NADP(+) adjacent to the FAD molecule, suitable for electron transfer, without any readjustments of the conformation of NADP(+). A comparison of the binding surfaces of thioredoxin and thioredoxin reductases from H. pylori and E. coli shows that the overall surface charge distribution in these proteins is conserved and that residue substitutions that change the shape of the binding surface may account for the species-specific recognition of thioredoxin by TrxR.
- The Medical Nobel Institute for Biochemistry, Department of Medical Biochemistry and Biophysics, Karolinska Institutet, S-171 77 Stockholm, Sweden.
Organizational Affiliation: