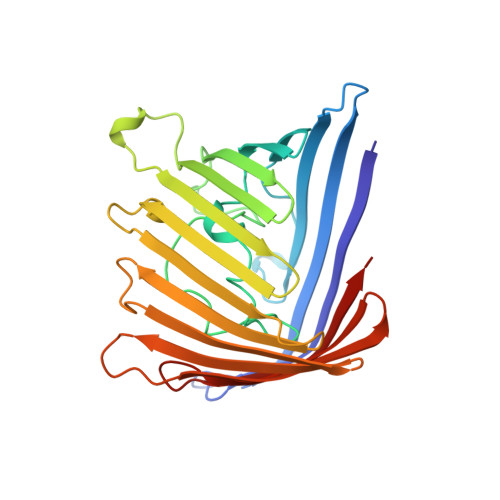
Porin mutants with new channel properties.
Schmid, B., Maveyraud, L., Kromer, M., Schulz, G.E.(1998) Protein Sci 7: 1603-1611
- PubMed: 9684893 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.5560070714
- Primary Citation Related Structures:
1BH3, 2PRN, 3PRN, 5PRN, 6PRN, 7PRN, 8PRN - PubMed Abstract:
The general diffusion porin from Rhodopseudomonas blastica was produced in large amounts in Escherichia coli inclusion bodies and (re)natured to the exact native structure. Here, we report on 13 mutants at the pore eyelet giving rise to new diffusion properties as measured in planar lipid bilayer experiments. The crystal structures of seven of these mutants were established. The effects of charge-modifying mutations at the pore eyelet are consistent with the known selectivity for cations. Deletions of 16 and 27 residues of the constriction loop L3 resulted in labile trimers and pores. The reduction of the eyelet cross section by introducing tryptophans gave rise to a closely correlated decrease of the conductivities. A mutant with six newly introduced tryptophans in the eyelet closed its pore in a defined manner within seconds under a voltage of 20 mV, suggesting the existence of two states. The results indicate that the pore can be engineered in a rational manner.
- Institut für Organische Chemie und Biochemie, Albert-Ludwigs-Universität, Freiburg im Breisgau, Germany.
Organizational Affiliation: