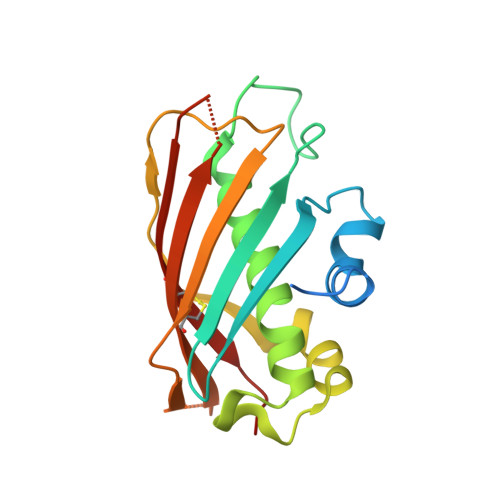
Identification of a type III thioesterase reveals the function of an operon crucial for Mtb virulence.
Wang, F., Langley, R., Gulten, G., Wang, L., Sacchettini, J.C.(2007) Chem Biol 14: 543-551
- PubMed: 17524985 Search on PubMed
- DOI: https://doi.org/10.1016/j.chembiol.2007.04.005
- Primary Citation Related Structures:
2PFC - PubMed Abstract:
Rv0098 is part of an operon, Rv0096-Rv0101, from Mycobacterium tuberculosis (Mtb) that is essential for Mtb's survival in mouse macrophages. This operon also contains an acyl carrier protein and one of the only two nonribosomal peptide synthases in Mtb. Rv0098 is annotated in the genome as a hypothetical protein and was proposed to be an acyl-coenzyme A (CoA) dehydratase. The structure of Rv0098, together with subsequent biochemical analysis, indicated that Rv0098 is a long-chain fatty acyl-CoA thioesterase (FcoT). However, FcoT lacks a general base or a nucleophile that is always found in the catalytic site of type II and type I thioesterases, respectively. The active site of Mtb FcoT reveals the structural basis for its substrate specificity for long-chain acyl-CoA and allows us to propose a catalytic mechanism for the enzyme. The characterization of Mtb FcoT provides a putative function of this operon that is crucial for Mtb pathogenicity.
- Department of Biochemistry and Biophysics, Texas A&M University, College Station, TX 77843, USA.
Organizational Affiliation: