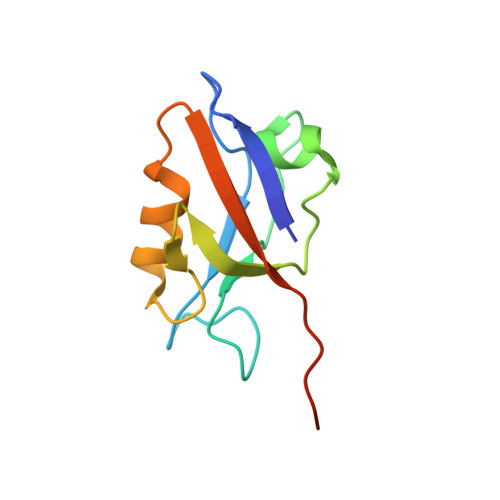
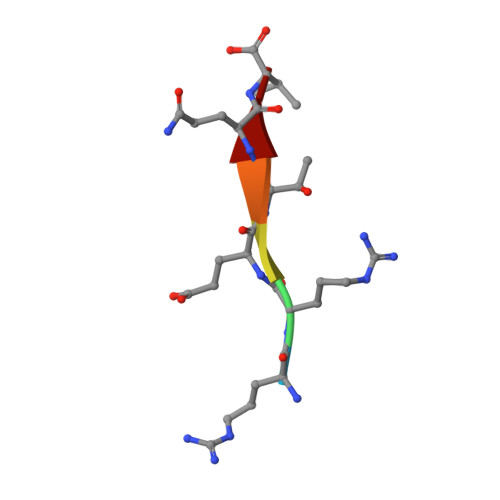
Solution structure of the hDlg/SAP97 PDZ2 domain and its mechanism of interaction with HPV-18 papillomavirus E6 protein.
Liu, Y., Henry, G.D., Hegde, R.S., Baleja, J.D.(2007) Biochemistry 46: 10864-10874
- PubMed: 17713926 Search on PubMed
- DOI: https://doi.org/10.1021/bi700879k
- Primary Citation Related Structures:
2OQS - PubMed Abstract:
The E6 protein from high-risk types of human papillomavirus (HPV) binds PDZ-domain containing proteins and targets them for degradation. We used isothermal titration calorimetry to measure the interaction of a peptide from the C-terminus of HPV-18 E6 to the second PDZ domain (PDZ2) from the human homologue of the Drosophila discs large tumor suppressor protein (hDlg). Isothermal titration calorimetry experiments with a series of peptides showed that HPV-18 E6 bound hDlg PDZ2 about 5-fold stronger than HPV-16 E6, that the contribution of Arg154 to binding was about 1 kcal/mol, and that the binding was disabled by phosphorylation at Thr156. We then used NMR to determine the solution structure of the complex of PDZ2 bound to the HPV-18 E6 peptide. The resultant structures were of high quality and had backbone root-mean-square deviations of less than 0.5 A. The structure shows a novel mode of interaction in which six residues of the HPV-18 E6 peptide are contacted by the PDZ2 domain, in contrast to the typical four residues used by class I PDZ domains. Molecular dynamics simulations supported a model in which the C- and N-terminal ends of the peptide had different mobilities within the complex. Comparison of the NMR complex structure to previously determined X-ray structures of PDZ2 by itself and bound to different peptides allows a description of conformational changes required for PDZ2 to bind to HPV-18 E6.
- Department of Biochemistry, Tufts University School of Medicine, 136 Harrison Avenue, Boston, Massachusetts 02111, USA.
Organizational Affiliation: