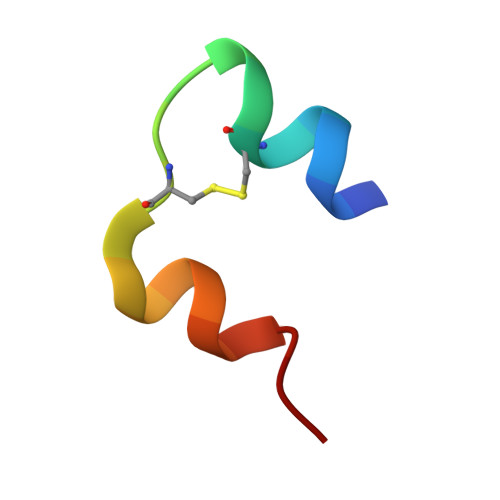
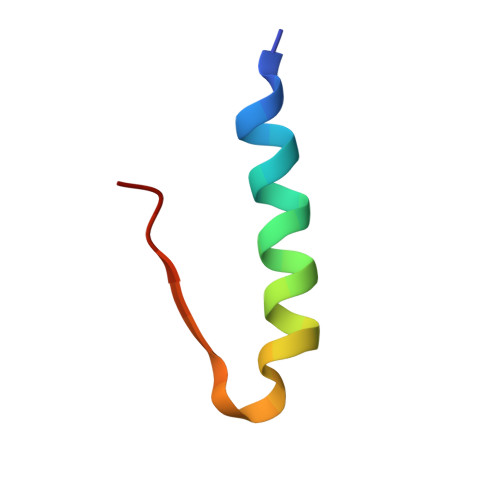
Crystallographic characterization of two novel crystal forms of human insulin induced by chaotropic agents and a shift in pH.
Norrman, M., Schluckebier, G.(2007) BMC Struct Biol 7: 83-83
- PubMed: 18093308 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1186/1472-6807-7-83
- Primary Citation Related Structures:
2OLY, 2OLZ, 2OM0, 2OM1 - PubMed Abstract:
Insulin is a therapeutic protein that is widely used for the treatment of diabetes. Its biological function was discovered more than 80 years ago and it has since then been characterized extensively. Crystallization of the insulin molecule has always been a key activity since the protein is often administered by subcutaneous injections of crystalline insulin formulations. Over the years, insulin has been crystallized and characterized in a number of crystal systems. Interestingly, we have now discovered two new crystal forms of human insulin. The crystals were obtained when the two chaotropic agents, urea and thiocyanate were present in the crystallization experiments, and their structures were determined by X-ray crystallography. The crystals belong to the orthorhombic and monoclinic crystal systems, with space groups C2221 and C2 respectively. The orthorhombic crystals were obtained at pH 6.5 and contained three insulin hexamers in R6 conformation in the asymmetric unit whilst the monoclinic C2 crystals were obtained at pH 7.0 and contained one R6 hexamer in the asymmetric unit. Common for the two new crystals is a hexamer-hexamer interaction that has not been found in any of the previous crystal forms of insulin. The contacts involve a tight glutamate-glutamate interaction with a distance of 2.3 A between groups. The short distance suggests a low barrier hydrogen bond. In addition, two tyrosine-tyrosine interactions occupying a known phenol binding pocket contribute to the stabilization of the contacts. Within the crystals, distinct binding sites for urea were found, adding further to the discussion on the role of urea in protein denaturation. The change in space group from C2221 to C2 was primarily caused by an increase in pH. The fewer number of hexamer-hexamer interactions comprising the short hydrogen bond in the C2 space group suggest that pH is the driving force. In addition, the distance between the two glutamates increases from 2.32 A in the C2221 crystals to 2.4 A in the C2 crystals. However, in both cases the low barrier hydrogen bond and the tyrosine-tyrosine interaction should contribute to the stability of the crystals which is crucial when used in pharmaceutical formulations.
- Diabetes Protein Engineering, Novo Nordisk A/S, Novo Nordisk Park, DK-2760 Måløv, Denmark. mtno@novonordisk.com
Organizational Affiliation: