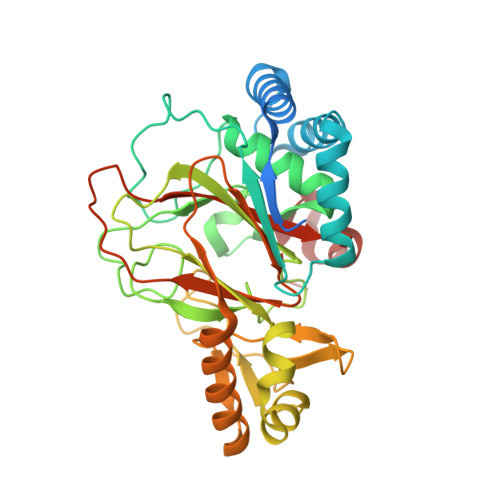
Mechanistic and structural basis of stereospecific Cbeta-hydroxylation in calcium-dependent antibiotic, a daptomycin-type lipopeptide
Strieker, M., Kopp, F., Mahlert, C., Essen, L.-O., Marahiel, M.A.(2007) ACS Chem Biol 2: 187-196
- PubMed: 17373765 Search on PubMed
- DOI: https://doi.org/10.1021/cb700012y
- PubMed Abstract:
Non-ribosomally synthesized lipopeptide antibiotics of the daptomycin type are known to contain unnatural beta-modified amino acids, which are essential for bioactivity. Here we present the biochemical and structural basis for the incorporation of 3-hydroxyasparagine at position 9 in the 11-residue acidic lipopeptide lactone calcium-dependent antibiotic (CDA). Direct hydroxylation of l-asparagine by AsnO, a non-heme Fe(2+)/alpha-ketoglutarate-dependent oxygenase encoded by the CDA biosynthesis gene cluster, was validated by Fmoc derivatization of the reaction product and LC/MS analysis. The 1.45, 1.92, and 1.66 A crystal structures of AsnO as apoprotein, Fe(2+) complex, and product complex, respectively, with (2S,3S)-3-hydroxyasparagine and succinate revealed the stereoselectivity and substrate specificity of AsnO. The comparison of native and product-complex structures of AsnO showed a lid-like region (residues F208-E223) that seals the active site upon substrate binding and shields it from sterically demanding peptide substrates. Accordingly, beta-hydroxylated asparagine is synthesized prior to its incorporation into the growing CDA peptide. The AsnO structure could serve as a template for engineering novel enzymes for the synthesis of beta-hydroxylated amino acids.
- Department of Chemistry/Biochemistry, Philipps University of Marburg, Marburg, Germany.
Organizational Affiliation: