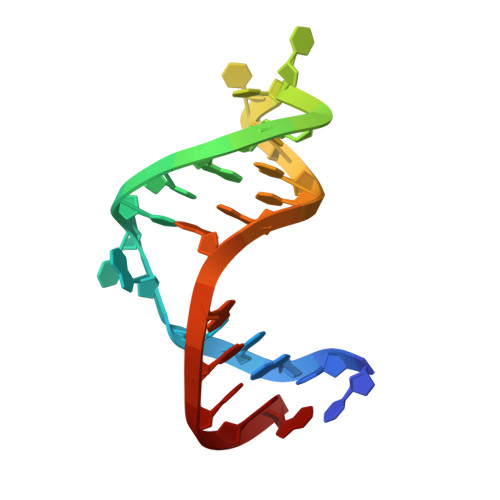
Solution Structure of a Designed Spirocyclic Helical Ligand Binding at a Two-Base Bulge Site in DNA.
Zhang, N., Lin, Y., Xiao, Z., Jones, G.B., Goldberg, I.H.(2007) Biochemistry 46: 4793-4803
- PubMed: 17388570 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi602599d
- Primary Citation Related Structures:
2OEY - PubMed Abstract:
The solution structure of the complex formed between an oligodeoxynucleotide containing a two-base bulge (5'-CCATCGTCTACCTTTGGTAGGATGG) and SCA-alpha2, a designed spirocyclic helical molecule, has been elucidated. SCA-alpha2, a close mimic of the metabolite, NCSi-gb, of the DNA bulge-specific enediyne antibiotic neocarzinostatin, differs in possessing a more stable spirocyclic ring system and in lacking certain bulky groupings that compromise bulged DNA binding. This study provides a detailed comparison of the binding modes of the two complexes and provides new insights into the importance of shape and space, as opposed to simple nucleotide sequence, in complex formation at the bulge site. The two rigidly held aromatic rings of SCA-alpha2 form a right-handed helical molecular wedge that specifically penetrates the bulge-binding pocket and immobilizes the two bulge residues (GT), which point toward the minor groove, rather than the major groove as in the NCSi-gb.bulged DNA complex. The ligand aromatic ring systems stack on the DNA bulge-flanking base pairs that define the long sides of the triangular prism binding pocket. Like NCSi-gb, SCA-alpha2 possesses the natural N-methylfuranose moiety, alpha-linked to the benzindanol (BI) moiety. The amino sugar anchors in the major groove of the DNA and points toward the 3'-bulge-flanking base pair. Lacking the bulky cyclocarbonate of NCSi-gb, the SCA-alpha2.bulged DNA complex has a much less twisted and buckled 3'-bulge-flanking base pair (dG20.dC8), and the G20 residue stacks directly above the BI ring platform. Also, the absence of the methyl group and the free rotation of the methoxy group on the dihydronaphthanone (NA) moiety of SCA-alpha2 allow better stacking geometry of the NA ring above the 5'-bulge-flanking dG21.dC5 base pair. These and other considerations help to explain why NCSi-gb binds very poorly to bulged RNA and are consistent with the recent observation of good binding with SCA-alpha2. Thus, although the two complexes resemble each other closely, they differ in important local environmental details. SCA-alpha2 has a better hand-in-glove fit at the bulge site, making it an ideal platform for the placement of moieties that can react covalently with the DNA and for generating congeners specific for bulges in RNA.
- Department of Biological Chemistry and Molecular Pharmacology, Harvard Medical School, Boston, Massachusetts 02115, USA.
Organizational Affiliation: