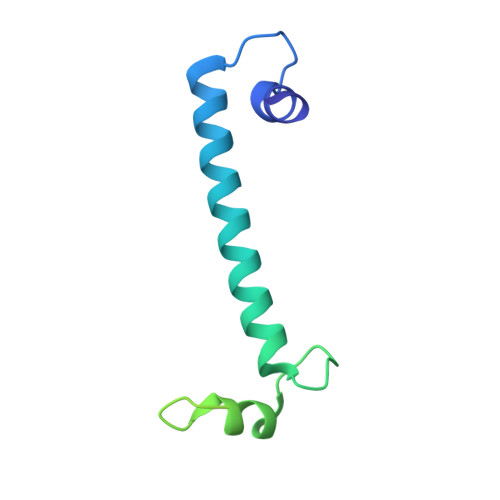
Crystal structure of calcium binding protein-1 from Entamoeba histolytica: A novel arrangement of EF hand motifs.
Kumar, S., Padhan, N., Alam, N., Gourinath, S.(2007) Proteins 68: 990-998
- PubMed: 17554780 Search on PubMed
- DOI: https://doi.org/10.1002/prot.21455
- Primary Citation Related Structures:
2NXQ, 5XOP - PubMed Abstract:
Calcium plays a pivotal role in the pathogenesis of amoebiasis, a major disease caused by Entamoeba histolytica. Several EF-hand containing calcium-binding proteins (CaBPs) have been identified from E. histolytica. Even though these proteins have very high sequence similarity, they bind to different target proteins in a Ca2+ dependent manner, leading to different functional pathways (Yadava et al., Mol Biochem Parasito 1997;84:69-82; Chakrabarty et al., J Biol Chem 2004;279:12898-12908) The crystal structure of the Entamoeba histolytica calcium binding protein-1 (EhCaBP1) has been determined at 2.4 A resolution. The crystals were grown using MPD as precipitant and they belong to P6(3) space group with unit cell parameters of a = 95.25 A, b = 95.25 A, c = 64.99 A. Only two out of the four expected EF hand motifs could be modeled into the electron density map and the final model refined to R factor of 25.6% and Free_R of 28%. Unlike CaM, the first two EF hand motifs in EhCaBP1 are connected by a long helix and form a dumbbell shaped structure. Owing to domain swapping oligomerization three EhCaBP1 molecules interact in a head to tail manner to form a triangular trimer. This arrangement allows the EF-hand motif of one molecule to interact with that of an adjacent molecule to form a two EF-hand domain similar to that seen in the N-terminal domain of the NMR structure of CaBP1, calmodulin and troponin C. The oligomeric state of EhCaBP1 results in reduced flexibility between domains and may be responsible for the more limited set of targets recognized by EhCaBP1.
- School of Life Sciences, Jawaharlal Nehru University, New Delhi 110067, India.
Organizational Affiliation: