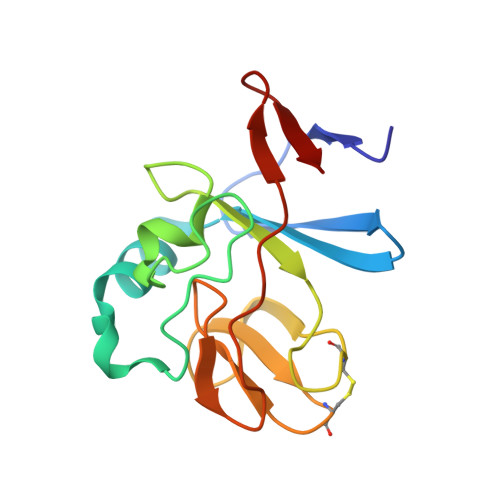
Atomic resolution structures of rieske iron-sulfur protein: role of hydrogen bonds in tuning the redox potential of iron-sulfur clusters.
Kolling, D.J., Brunzelle, J.S., Lhee, S., Crofts, A.R., Nair, S.K.(2007) Structure 15: 29-38
- PubMed: 17223530 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2006.11.012
- Primary Citation Related Structures:
2NUK, 2NUM, 2NVE, 2NVF, 2NVG, 2NWF - PubMed Abstract:
The Rieske [2Fe-2S] iron-sulfur protein of cytochrome bc(1) functions as the initial electron acceptor in the rate-limiting step of the catalytic reaction. Prior studies have established roles for a number of conserved residues that hydrogen bond to ligands of the [2Fe-2S] cluster. We have constructed site-specific variants at two of these residues, measured their thermodynamic and functional properties, and determined atomic resolution X-ray crystal structures for the native protein at 1.2 A resolution and for five variants (Ser-154-->Ala, Ser-154-->Thr, Ser-154-->Cys, Tyr-156-->Phe, and Tyr-156-->Trp) to resolutions between 1.5 A and 1.1 A. These structures and complementary biophysical data provide a molecular framework for understanding the role hydrogen bonds to the cluster play in tuning thermodynamic properties, and hence the rate of this bioenergetic reaction. These studies provide a detailed structure-function dissection of the role of hydrogen bonds in tuning the redox potentials of [2Fe-2S] clusters.
- Center for Biophysics and Computational Biology, University of Illinois at Urbana-Champaign, 600 S. Mathews Avenue, Urbana, IL 61801, USA.
Organizational Affiliation: