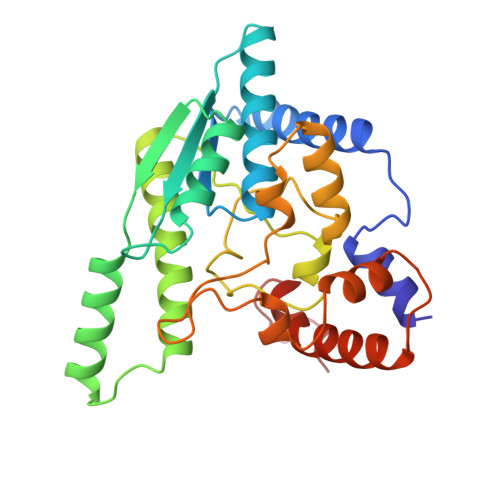
A novel deamido-NAD+-binding site revealed by the trapped NAD-adenylate intermediate in the NAD+ synthetase structure.
Rizzi, M., Bolognesi, M., Coda, A.(1998) Structure 6: 1129-1140
- PubMed: 9753692 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(98)00114-2
- Primary Citation Related Structures:
2NSY - PubMed Abstract:
Nicotinamide adenine dinucleotide (NAD+) has a central role in life processes. The ubiquitous enzyme NAD+ synthetase catalyzes a key step in NAD+ biosynthesis, transforming deamido-NAD+ into NAD+ by a two-step reaction. NAD+ synthetase belongs to the amidotransferase family and has been recognized as a member of the family of N-type ATP pyrophosphatases. In order to investigate the mechanism of the reaction carried out by NAD+ synthetase we have determined a high-resolution three-dimensional structure of the Bacillus subtilis homodimeric NAD+ synthetase in complex with the trapped reaction intermediate NAD-adenylate. Two NAD-adenylate molecules and two pyrophosphate (PPi) molecules are observed in the 1.3 A resolution structure of the NAD+ synthetase-NAD-adenylate complex. Structural studies on the NAD+ synthetase-NAD-adenylate adduct and on the cation-binding sites reveal a new deamido-NAD+-binding site located at the subunit interface, locate a binuclear magnesium cluster at the ATP-binding site and, identify two monovalent cation sites, one of which may represent an ammonium-binding site. Our results suggest that two different catalytic strategies have been adopted by NAD+ synthetase in the two different steps of the reaction. During the adenylation step, no protein residues seem to be located properly to directly participate in catalysis, which is likely to be carried out with the fundamental assistance of an electron-withdrawing trimetallic constellation present in the active site. A different behavior is observed for the second step, in which an ammonium ion is the binding species. In this step, Asp173 is a key residue in both deprotonation of the primarily bound ammonium ion, and stabilization of the tetrahedral transition-state intermediate. Moreover, the structural data suggest that product release can take place only after all substrates are bound to the enzyme, and product release is ultimately controlled by the conformation adopted by two mobile loops.
- Department of Pharmaceutical Science and Technology University of Torino, Italy. rizzi@ipvgen.unipv.it
Organizational Affiliation: