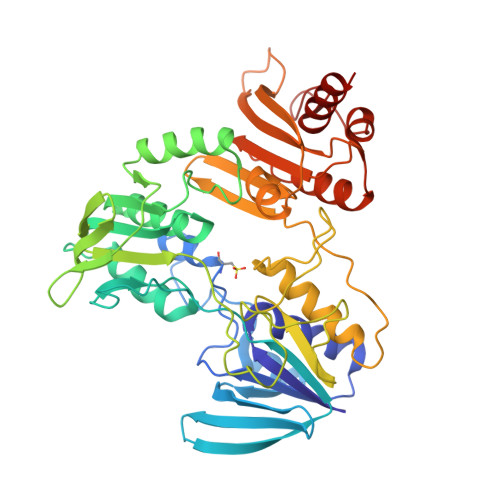
NADH binding site and catalysis of NADH peroxidase.
Stehle, T., Claiborne, A., Schulz, G.E.(1993) Eur J Biochem 211: 221-226
- PubMed: 8425532 Search on PubMed
- DOI: https://doi.org/10.1111/j.1432-1033.1993.tb19889.x
- Primary Citation Related Structures:
2NPX - PubMed Abstract:
The structure of the complex between cofactor NADH and the enzyme NADH peroxidase from Streptococcus faecalis 10C1 (Enterococcus faecalis) has been determined by crystal soaking, X-ray data collection, model building of NADH and refinement at 0.24-nm resolution based on the known enzyme structure [Stehle, T., Ahmed, S. A., Claiborne, A. & Schulz, G. E. (1991) J. Mol. Biol. 221, 1325-1344]. Apart from NADH, the catalytic center of the enzyme contains FAD and a cysteine that shuttles between thiolate and sulfenic acid states. Unfortunately, this cysteine was irreversibly oxidized to a cysteine sulfonic acid in the established enzyme structure. Based on the geometry of the catalytic center, we discuss the stabilization of the oxidation-sensitive sulfenic acid and propose a reaction mechanism.
- Institut für Organische Chemie und Biochemie der Universität, Freiburg im Breisgau, Federal Republic of Germany.
Organizational Affiliation: