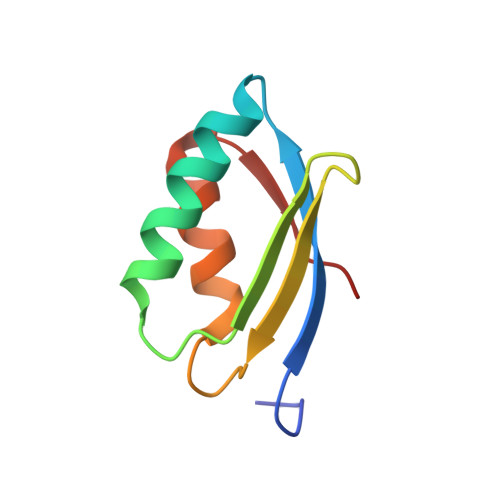
The Structure of Metal Binding Domain 1 of the Copper Transporter ATP7B Reveals Mechanism of a Singular Wilson Disease Mutation.
Yu, C.H., Lee, W., Nokhrin, S., Dmitriev, O.Y.(2018) Sci Rep 8: 581-581
- PubMed: 29330485 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-017-18951-1
- Primary Citation Related Structures:
2N7Y - PubMed Abstract:
Copper-transporter ATP7B maintains copper homeostasis in the human cells and delivers copper to the biosynthetic pathways for incorporation into the newly synthesized copper-containing proteins. ATP7B is a target of several hundred mutations that lead to Wilson disease, a chronic copper toxicosis. ATP7B contains a chain of six cytosolic metal-binding domains (MBDs), the first four of which (MBD1-4) are believed to be regulatory, and the last two (MBD5-6) are required for enzyme activity. We report the NMR structure of MBD1, the last unsolved metal-binding domain of ATP7B. The structure reveals the disruptive mechanism of G85V mutation, one of the very few disease causing missense mutations in the MBD1-4 region of ATP7B.
- Department of Biochemistry, University of Saskatchewan, Saskatoon, SK, Canada.
Organizational Affiliation: