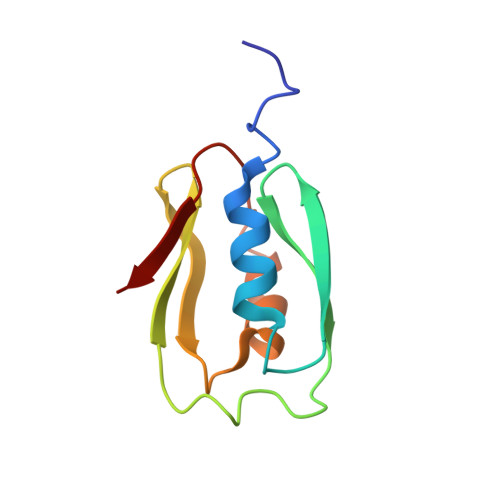
NMR structure of the Bacillus cereus hemolysin II C-terminal domain reveals a novel fold.
Kaplan, A.R., Kaus, K., De, S., Olson, R., Alexandrescu, A.T.(2017) Sci Rep 7: 3277-3277
- PubMed: 28607368 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-017-02917-4
- Primary Citation Related Structures:
2N67 - PubMed Abstract:
In addition to multiple virulence factors, Bacillus cereus a pathogen that causes food poisoning and life-threatening wound infections, secretes the pore-forming toxin hemolysin II (HlyII). The HlyII toxin has a unique 94 amino acid C-terminal domain (HlyIIC). HlyIIC exhibits splitting of NMR resonances due to cis/trans isomerization of a single proline near the C-terminus. To overcome heterogeneity, we solved the structure of P405M-HlyIIC, a mutant that exclusively stabilizes the trans state. The NMR structure of HlyIIC reveals a novel fold, consisting of two subdomains αA-β1-β2 and β3-β4-αB-β5, that come together in a barrel-like structure. The barrel core is fastened by three layers of hydrophobic residues. The barrel end opposite the HlyIIC-core has a positively charged surface, that by binding negatively charged moieties on cellular membranes, may play a role in target-cell surface recognition or stabilization of the heptameric pore complex. In the WT domain, dynamic flexibility occurs at the N-terminus and the first α-helix that connects the HlyIIC domain to the HlyII-core structure. In the destabilizing P405M mutant, increased flexibility is evident throughout the first subdomain, suggesting that the HlyIIC structure may have arisen through gene fusion.
- Department of Molecular and Cell Biology, University of Connecticut, 91 N. Eagleville Rd, Storrs, CT, 06269-3125, USA.
Organizational Affiliation: