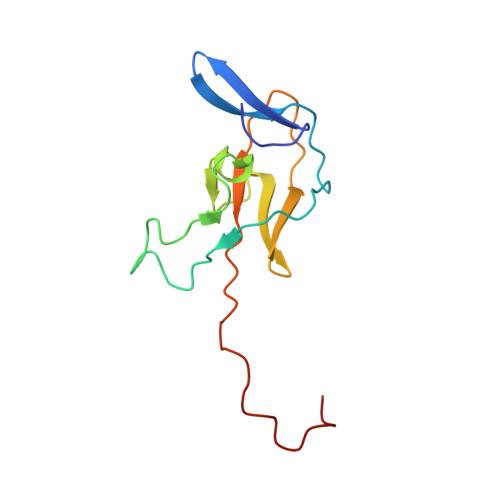
The Identification and Structure of an N-Terminal PR Domain Show that FOG1 Is a Member of the PRDM Family of Proteins.
Clifton, M.K., Westman, B.J., Thong, S.Y., O'Connell, M.R., Webster, M.W., Shepherd, N.E., Quinlan, K.G., Crossley, M., Blobel, G.A., Mackay, J.P.(2014) PLoS One 9: e106011-e106011
- PubMed: 25162672 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0106011
- Primary Citation Related Structures:
2MPL - PubMed Abstract:
FOG1 is a transcriptional regulator that acts in concert with the hematopoietic master regulator GATA1 to coordinate the differentiation of platelets and erythrocytes. Despite considerable effort, however, the mechanisms through which FOG1 regulates gene expression are only partially understood. Here we report the discovery of a previously unrecognized domain in FOG1: a PR (PRD-BF1 and RIZ) domain that is distantly related in sequence to the SET domains that are found in many histone methyltransferases. We have used NMR spectroscopy to determine the solution structure of this domain, revealing that the domain shares close structural similarity with SET domains. Titration with S-adenosyl-L-homocysteine, the cofactor product synonymous with SET domain methyltransferase activity, indicated that the FOG PR domain is not, however, likely to function as a methyltransferase in the same fashion. We also sought to define the function of this domain using both pulldown experiments and gel shift assays. However, neither pulldowns from mammalian nuclear extracts nor yeast two-hybrid assays reproducibly revealed binding partners, and we were unable to detect nucleic-acid-binding activity in this domain using our high-diversity Pentaprobe oligonucleotides. Overall, our data demonstrate that FOG1 is a member of the PRDM (PR domain containing proteins, with zinc fingers) family of transcriptional regulators. The function of many PR domains, however, remains somewhat enigmatic for the time being.
- School of Molecular Bioscience, University of Sydney, Sydney, NSW, Australia.
Organizational Affiliation: