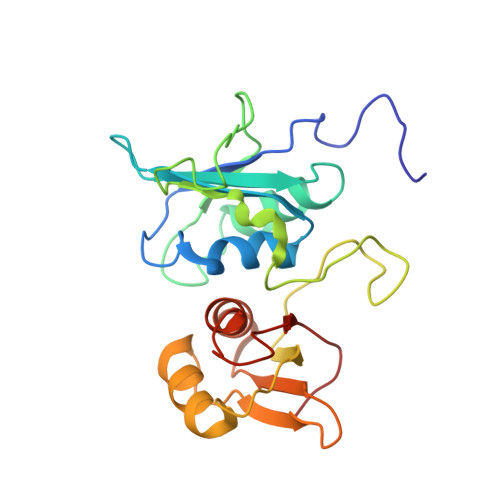
Solution and crystal structures of a C-terminal fragment of the neuronal isoform of the polypyrimidine tract binding protein (nPTB).
Joshi, A., Esteve, V., Buckroyd, A.N., Blatter, M., Allain, F.H., Curry, S.(2014) PeerJ 2: e305-e305
- PubMed: 24688880 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7717/peerj.305
- Primary Citation Related Structures:
2MJU, 4CQ1 - PubMed Abstract:
The eukaryotic polypyrimidine tract binding protein (PTB) serves primarily as a regulator of alternative splicing of messenger RNA, but is also co-opted to other roles such as RNA localisation and translation initiation from internal ribosome entry sites. The neuronal paralogue of PTB (nPTB) is 75% identical in amino acid sequence with PTB. Although the two proteins have broadly similar RNA binding specificities and effects on RNA splicing, differential expression of PTB and nPTB can lead to the generation of alternatively spliced mRNAs. RNA binding by PTB and nPTB is mediated by four RNA recognition motifs (RRMs). We present here the crystal and solution structures of the C-terminal domain of nPTB (nPTB34) which contains RRMs 3 and 4. As expected the structures are similar to each other and to the solution structure of the equivalent fragment from PTB (PTB34). The result confirms that, as found for PTB, RRMs 3 and 4 of nPTB interact with one another to form a stable unit that presents the RNA-binding surfaces of the component RRMs on opposite sides that face away from each other. The major differences between PTB34 and nPTB34 arise from amino acid side chain substitutions on the exposed β-sheet surfaces and adjoining loops of each RRM, which are likely to modulate interactions with RNA.
- Department of Life Sciences, Imperial College, London, United Kingdom.
Organizational Affiliation: