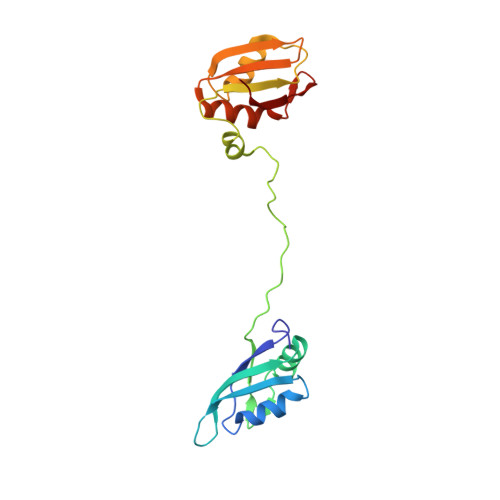
Structure, dynamics and RNA binding of the multi-domain splicing factor TIA-1.
Wang, I., Hennig, J., Jagtap, P.K., Sonntag, M., Valcarcel, J., Sattler, M.(2014) Nucleic Acids Res 42: 5949-5966
- PubMed: 24682828 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gku193
- Primary Citation Related Structures:
2MJN - PubMed Abstract:
Alternative pre-messenger ribonucleic acid (pre-mRNA) splicing is an essential process in eukaryotic gene regulation. The T-cell intracellular antigen-1 (TIA-1) is an apoptosis-promoting factor that modulates alternative splicing of transcripts, including the pre-mRNA encoding the membrane receptor Fas. TIA-1 is a multi-domain ribonucleic acid (RNA) binding protein that recognizes poly-uridine tract RNA sequences to facilitate 5' splice site recognition by the U1 small nuclear ribonucleoprotein (snRNP). Here, we characterize the RNA interaction and conformational dynamics of TIA-1 by nuclear magnetic resonance (NMR), isothermal titration calorimetry (ITC) and small angle X-ray scattering (SAXS). Our NMR-derived solution structure of TIA-1 RRM2-RRM3 (RRM2,3) reveals that RRM2 adopts a canonical RNA recognition motif (RRM) fold, while RRM3 is preceded by an non-canonical helix α0. NMR and SAXS data show that all three RRMs are largely independent structural modules in the absence of RNA, while RNA binding induces a compact arrangement. RRM2,3 binds to pyrimidine-rich FAS pre-mRNA or poly-uridine (U9) RNA with nanomolar affinities. RRM1 has little intrinsic RNA binding affinity and does not strongly contribute to RNA binding in the context of RRM1,2,3. Our data unravel the role of binding avidity and the contributions of the TIA-1 RRMs for recognition of pyrimidine-rich RNAs.
- Institute of Structural Biology, Helmholtz Zentrum München, Ingolstädter Landstraße 1, 85764 Neuherberg, Germany Center for Integrated Protein Science Munich and Biomolecular NMR, Department Chemie Technische Universität München, Lichtenbergstraße 4, 85747 Garching, Germany.
Organizational Affiliation: