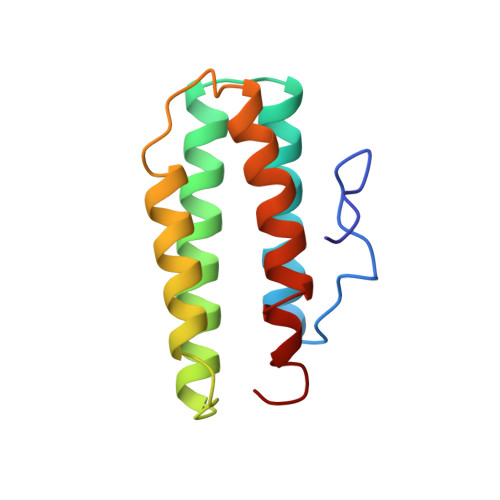
Structure of myohemerythrin in the azidomet state at 1.7/1.3 A resolution.
Sheriff, S., Hendrickson, W.A., Smith, J.L.(1987) J Mol Biology 197: 273-296
- PubMed: 3681996 Search on PubMed
- DOI: https://doi.org/10.1016/0022-2836(87)90124-0
- Primary Citation Related Structures:
2MHR - PubMed Abstract:
The molecular model of myohemerythrin, an oxygen-carrying protein from sipunculan worms, has been refined by stereochemically restrained least-squares minimization at 1.7/1.3 A resolution to a conventional R-value of 0.158. The estimated positional standard deviation is better than 0.15 A for most of the 979 protein atoms. The average isotropic displacement parameter, B, for the protein atoms is 23.1 A2. This high average B parameter appears to be due to the overall motion of the molecule, which correlates with the observed anisotropic diffraction. The side-chains of seven residues were modeled in two conformations, i.e. the side-chains were discretely disordered, and B parameters for several lysine and glutamate side-chains indicate that they are poorly localized. Of the residues in myohemerythrin, 66% are helical, with 62% occurring in four long alpha-helices with mean values for the backbone torsion angles of phi = -65 degrees, psi = -42 degrees, and for the hydrogen bonds distances of N ... O, 3.0 A and H ... O, 2.1 A, and angles of N ... O = C, 153 degrees, N-H ... O, 157 degrees, and H ... O = C, 147 degrees. For two-thirds of the alpha-helical residues, the torsional rotation of the C alpha-C beta bond, chi 1, is approximately -60 degrees, and for one-third chi 1 is approximately 180 degrees. Although most turns in myohemerythrin are well-categorized by previous classification, two do not fit in established patterns. Also included in the refined model are three sulfate ions, all partially occupied, and 157 water molecules, 40% of which are modeled fully occupied. Only one water molecule is internal to the protein, the remainder occur on the surface and are observed principally between symmetry-related molecules contributing, along with van der Waals' contacts, most of the interactions between molecules. There are eight intermolecular protein-protein hydrogen bonds, of which only four are between well-located atoms.
- Department of Biochemistry and Molecular Biophysics, Columbia University, New York, NY 10032.
Organizational Affiliation: