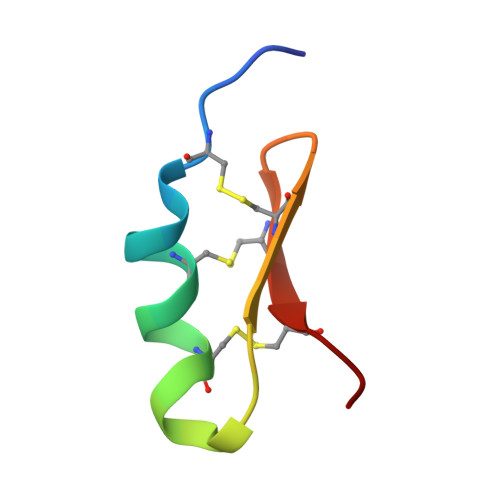
Cytotoxicity of recombinant tamapin and related toxin-like peptides on model cell lines.
Ramirez-Cordero, B., Toledano, Y., Cano-Sanchez, P., Hernandez-Lopez, R., Flores-Solis, D., Saucedo-Yanez, A.L., Chavez-Uribe, I., Brieba, L.G., Del Rio-Portilla, F.(2014) Chem Res Toxicol 27: 960-967
- PubMed: 24821061 Search on PubMed
- DOI: https://doi.org/10.1021/tx4004193
- Primary Citation Related Structures:
2KY3, 2LU9, 2ME7, 2MEL, 2MEN, 2MEO - PubMed Abstract:
The scorpion toxin tamapin displays the most potent and selective blockage against KCa2.2 channels known to date. In this work, we report the biosynthesis, three-dimensional structure, and cytotoxicity on cancer cell lines (Jurkat E6-1 and human mammary breast cancer MDA-MB-231) of recombinant tamapin and five related peptides bearing mutations on residues (R6A,R7A, R13A, R6A-R7A, and GS-tamapin) that were previously suggested to be important for tamapin's activity. The indicated cell lines were used as they constitutively express KCa2.2 channels. The studied toxin-like peptides displayed lethal responses on Jurkat T cells and breast cancer cells; their effect is dose- and time-dependent with IC50 values in the nanomolar range. The order of potency is r-tamapin>GS-tamapin>R6A>R13A>R6A-R7A>R7A for Jurkat T cells and r-tamapin>R7A for MDA-MB-231 breast cancer cells. Our structural determination by NMR demonstrated that r-tamapin preserves the folding of the αKTx5 subfamily and that neither single nor double alanine mutations affect the three-dimensional structure of the wild-type peptide. In contrast, our activity assays show that changes in cytotoxicity are related to the chemical nature of certain residues. Our results suggest that the toxic activity of r-tamapin on Jurkat and breast cancer cells could be mediated by the interaction of charged residues in tamapin with KCa2.2 channels via the apoptotic cell death pathway.
- Instituto de Química, Universidad Nacional Autónoma de México , Ciudad Universitaria, Circuito Exterior s/n, México, D.F. 04510, México.
Organizational Affiliation: