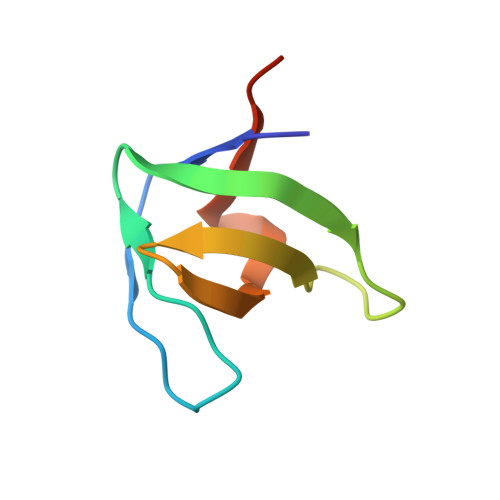
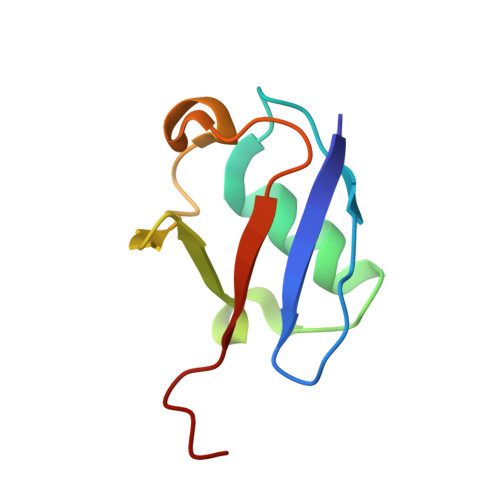
Distinct ubiquitin binding modes exhibited by SH3 domains: molecular determinants and functional implications.
Ortega Roldan, J.L., Casares, S., Ringkjbing Jensen, M., Cardenes, N., Bravo, J., Blackledge, M., Azuaga, A.I., van Nuland, N.A.(2013) PLoS One 8: e73018-e73018
- PubMed: 24039852 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0073018
- Primary Citation Related Structures:
2LZ6, 2MCN, 2YDL - PubMed Abstract:
SH3 domains constitute a new type of ubiquitin-binding domains. We previously showed that the third SH3 domain (SH3-C) of CD2AP binds ubiquitin in an alternative orientation. We have determined the structure of the complex between first CD2AP SH3 domain and ubiquitin and performed a structural and mutational analysis to decipher the determinants of the SH3-C binding mode to ubiquitin. We found that the Phe-to-Tyr mutation in CD2AP and in the homologous CIN85 SH3-C domain does not abrogate ubiquitin binding, in contrast to previous hypothesis and our findings for the first two CD2AP SH3 domains. The similar alternative binding mode of the SH3-C domains of these related adaptor proteins is characterised by a higher affinity to C-terminal extended ubiquitin molecules. We conclude that CD2AP/CIN85 SH3-C domain interaction with ubiquitin constitutes a new ubiquitin-binding mode involved in a different cellular function and thus changes the previously established mechanism of EGF-dependent CD2AP/CIN85 mono-ubiquitination.
- Departamento de Química Física e Instituto de Biotecnología, Facultad de Ciencias, Universidad de Granada, Granada, Spain ; Department of Biochemistry, University of Oxford, Oxford, United Kingdom.
Organizational Affiliation: