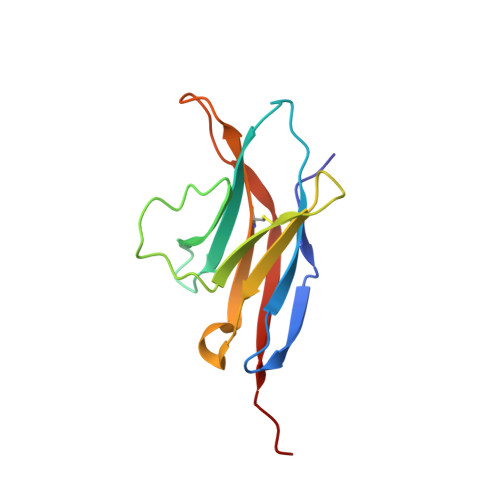
Structure and interactions of the human programmed cell death 1 receptor.
Cheng, X., Veverka, V., Radhakrishnan, A., Waters, L.C., Muskett, F.W., Morgan, S.H., Huo, J., Yu, C., Evans, E.J., Leslie, A.J., Griffiths, M., Stubberfield, C., Griffin, R., Henry, A.J., Jansson, A., Ladbury, J.E., Ikemizu, S., Carr, M.D., Davis, S.J.(2013) J Biological Chem 288: 11771-11785
- PubMed: 23417675 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M112.448126
- Primary Citation Related Structures:
2M2D - PubMed Abstract:
PD-1, a receptor expressed by T cells, B cells, and monocytes, is a potent regulator of immune responses and a promising therapeutic target. The structure and interactions of human PD-1 are, however, incompletely characterized. We present the solution nuclear magnetic resonance (NMR)-based structure of the human PD-1 extracellular region and detailed analyses of its interactions with its ligands, PD-L1 and PD-L2. PD-1 has typical immunoglobulin superfamily topology but differs at the edge of the GFCC' sheet, which is flexible and completely lacks a C" strand. Changes in PD-1 backbone NMR signals induced by ligand binding suggest that, whereas binding is centered on the GFCC' sheet, PD-1 is engaged by its two ligands differently and in ways incompletely explained by crystal structures of mouse PD-1 · ligand complexes. The affinities of these interactions and that of PD-L1 with the costimulatory protein B7-1, measured using surface plasmon resonance, are significantly weaker than expected. The 3-4-fold greater affinity of PD-L2 versus PD-L1 for human PD-1 is principally due to the 3-fold smaller dissociation rate for PD-L2 binding. Isothermal titration calorimetry revealed that the PD-1/PD-L1 interaction is entropically driven, whereas PD-1/PD-L2 binding has a large enthalpic component. Mathematical simulations based on the biophysical data and quantitative expression data suggest an unexpectedly limited contribution of PD-L2 to PD-1 ligation during interactions of activated T cells with antigen-presenting cells. These findings provide a rigorous structural and biophysical framework for interpreting the important functions of PD-1 and reveal that potent inhibitory signaling can be initiated by weakly interacting receptors.
- Radcliffe Department of Medicine, University of Oxford, John Radcliffe Hospital, Headington, Oxford OX3 9DU, United Kingdom.
Organizational Affiliation: