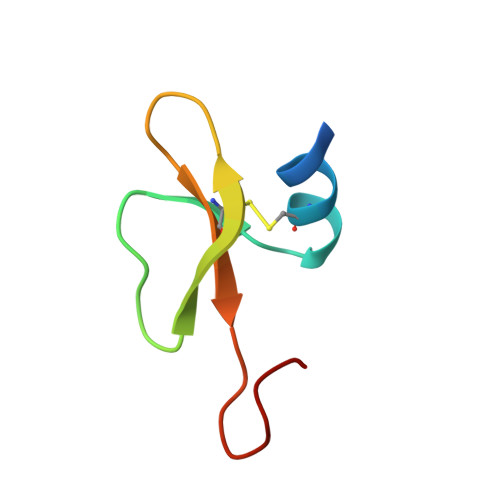
Structural Basis for the Interaction of Human beta-Defensin 6 and Its Putative Chemokine Receptor CCR2 and Breast Cancer Microvesicles.
De Paula, V.S., Gomes, N.S., Lima, L.G., Miyamoto, C.A., Monteiro, R.Q., Almeida, F.C., Valente, A.P.(2013) J Mol Biology 425: 4479-4495
- PubMed: 23938203 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2013.08.001
- Primary Citation Related Structures:
2LWL - PubMed Abstract:
Human β-defensins (hBDs) are believed to function as alarm molecules that stimulate the adaptive immune system when a threat is present. In addition to its antimicrobial activity, defensins present other activities such as chemoattraction of a range of different cell types to the sites of inflammation. We have solved the structure of the hBD6 by NMR spectroscopy that contains a conserved β-defensin domain followed by an extended C-terminus. We use NMR to monitor the interaction of hBD6 with microvesicles shed by breast cancer cell lines and with peptides derived from the extracellular domain of CC chemokine receptor 2 (Nt-CCR2) possessing or not possessing sulfation on Tyr26 and Tyr28. The NMR-derived model of the hBD6/CCR2 complex reveals a contiguous binding surface on hBD6, which comprises amino acid residues of the α-helix and β2-β3 loop. The microvesicle binding surface partially overlaps with the chemokine receptor interface. NMR spin relaxation suggests that free hBD6 and the hBD6/CCR2 complex exhibit microsecond-to-millisecond conformational dynamics encompassing the CCR2 binding site, which might facilitate selection of the molecular configuration optimal for binding. These data offer new insights into the structure-function relation of the hBD6-CCR2 interaction, which is a promising target for the design of novel anticancer agents.
- Centro Nacional de Ressonância Magnética Nuclear de Macromoléculas, Instituto de Bioquímica Médica, Universidade Federal do Rio de Janeiro, Rio de Janeiro, RJ 21941-902, Brazil.
Organizational Affiliation: