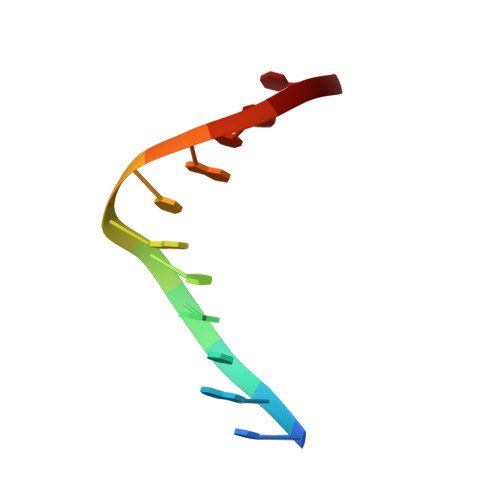
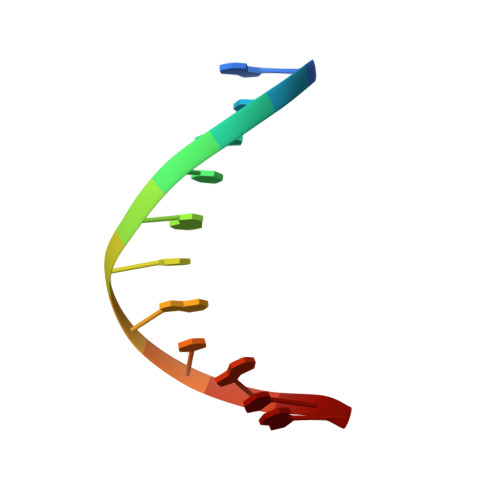
Structure and Stability of Duplex DNA Containing (5'S)-5',8-Cyclo-2'-deoxyadenosine: An Oxidatively Generated Lesion Repaired by NER.
Zaliznyak, T., Lukin, M., de Los Santos, C.(2012) Chem Res Toxicol 25: 2103-2111
- PubMed: 22928555 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/tx300193k
- Primary Citation Related Structures:
2LSF - PubMed Abstract:
Cellular respiration and ionizing radiation generate 5',8-cyclo-2'-deoxyribonucleosides, a special type of DNA damage that involves two modifications in the same nucleotide. These lesions evade the action of base excision glycosylases, and their removal is a function of the nucleotide excision repair pathway. Diastereomeric 5',8-cyclo-2'-deoxyadenosine blocks mammalian DNA replication, diminishes the levels of DNA transcription, and induces transcriptional mutagenesis. Using solution state NMR spectroscopy and restrained molecular dynamics simulations, we have determined the structure of an undecameric DNA duplex having a centrally located (5'S)-5',8-cyclo-2'-deoxyadenosine residue paired to T. The damaged duplex structure is a right-handed helix having Watson-Crick base-pair alignments throughout, and 2-deoxyribose puckers within the B-form conformation. Only small structural perturbations are observed at the lesion-containing and 5'-flanking base pair. The 2-deoxyribose of the damaged nucleotide adopts the O4'-exo conformation, and the S-cdA·T base pair is propeller twisted. The 5'-lesion-flanking base is tilted forming a significantly buckled base pair with its partner guanine. Analysis of UV-melting curves indicates mild thermal and thermodynamic destabilization on the damaged duplex. The S-cdA·T duplex structure shows many similarities to and some intriguing differences from the recently reported structure of an S-cdG·dC duplex³¹ that suggest different lesion site dynamics.
- Department of Pharmacological Sciences, School of Medicine, Stony Brook University, Stony Brook, New York 11794-8651, USA.
Organizational Affiliation: