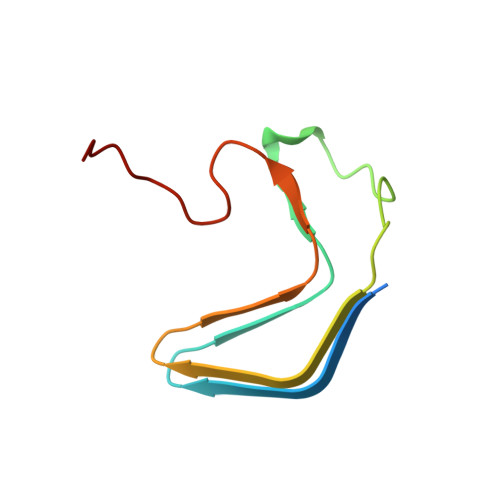
The Amyloid-Congo Red Interface at Atomic Resolution.
Schutz, A.K., Soragni, A., Hornemann, S., Aguzzi, A., Ernst, M., Bockmann, A., Meier, B.H.(2011) Angew Chem Int Ed Engl
- PubMed: 21591034 Search on PubMed
- DOI: https://doi.org/10.1002/anie.201008276
- Primary Citation Related Structures:
2LBU - Physikalische Chemie, ETH Zürich, 8093 Zürich, Switzerland.
Organizational Affiliation: