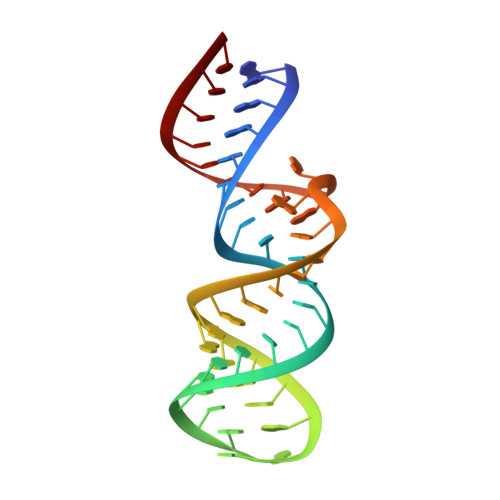
Structure of the HIV-1 Frameshift Site RNA Bound to a Small Molecule Inhibitor of Viral Replication.
Marcheschi, R.J., Tonelli, M., Kumar, A., Butcher, S.E.(2011) ACS Chem Biol 6: 857-864
- PubMed: 21648432 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/cb200082d
- Primary Citation Related Structures:
2L94 - PubMed Abstract:
Programmed -1 translational frameshifting is an essential event in the replication cycle of HIV. Frameshifting is required for expression of the viral Pol proteins, and drug-like molecules that target this process may inhibit HIV replication. A small molecule stimulator of HIV-1 frameshifting and inhibitor of viral replication, DB213 (RG501), was previously discovered from a high-throughput screen. However, the mechanistic basis for this compound's effects was unknown, and to date no structural information exists for small molecule effectors of frameshifting. Here, we investigate the binding of DB213 to the frameshift site RNA and have determined the structure of this complex by NMR. Binding of DB213 stabilizes the RNA and increases its melting temperature by 10 °C. The ligand binds to a primary site on the RNA stem-loop, although nonspecific interactions are also detected. The compound binds in the major groove and spans a distance of 9 base pairs. DB213 hydrogen bonds to phosphate groups on opposite sides of the major groove and alters the conformation of a conserved GGA bulge in the RNA. This study may provide a starting point for structure-based optimization of compounds targeting the HIV-1 frameshift site RNA.
- Department of Biochemistry, University of Wisconsin-Madison, Madison, Wisconsin 53706, United States.
Organizational Affiliation: