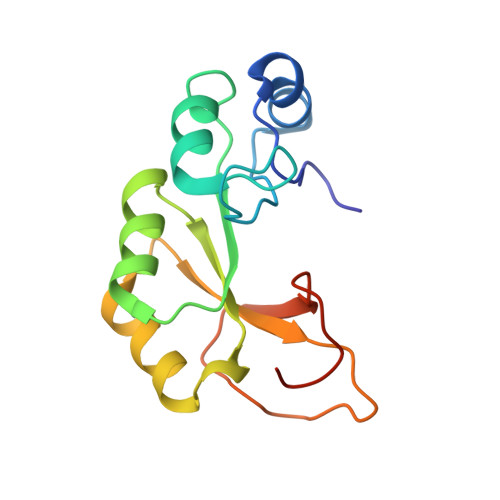
The Structure of a Truncated Phosphoribosylanthranilate Isomerase Suggests a Unified Model for Evolution of the (beta alpha)8 Barrel Fold
Setiyaputra, S., Mackay, J.P., Patrick, W.M.(2011) J Mol Biology 408: 291-303
- PubMed: 21354426 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2011.02.048
- Primary Citation Related Structures:
2KZH - PubMed Abstract:
The (βα)(8) barrel is one of the most common protein folds, and enzymes with this architecture display a remarkable range of catalytic activities. Many of these functions are associated with ancient metabolic pathways, and phylogenetic reconstructions suggest that the (βα)(8) barrel was one of the very first protein folds to emerge. Consequently, there is considerable interest in understanding the evolutionary processes that gave rise to this fold. In particular, much attention has been focused on the plausibility of (βα)(8) barrel evolution from homodimers of half barrels. However, we previously isolated a three-quarter-barrel-sized fragment of a (βα)(8) barrel, termed truncated phosphoribosylanthranilate isomerase (trPRAI), that is soluble and almost as thermostable as full-length N-(5'-phosphoribosyl)anthranilate isomerase (PRAI). Here, we report the NMR-derived structure of trPRAI. The subdomain is monomeric, is well ordered and adopts a native-like structure in solution. Side chains from strands β(1) (Glu3 and Lys5), β(2) (Tyr25) and β(6) (Lys122) of trPRAI repack to shield the hydrophobic core from the solvent. This result demonstrates that three-quarter barrels were viable intermediates in the evolution of the (βα)(8) barrel fold. We propose a unified model for (βα)(8) barrel evolution that combines our data, previously published work and plausible scenarios for the emergence of (initially error-prone) genetic systems. In this model, the earliest proto-cells contained diverse pools of part-barrel subdomains. Combinatorial assembly of these subdomains gave rise to many distinct lineages of (βα)(8) barrel proteins, that is, our model excludes the possibility that there was a single (βα)(8) barrel from which all present examples are descended.
- School of Molecular Bioscience, Darlington Campus, The University of Sydney, NSW 2006, Australia.
Organizational Affiliation: