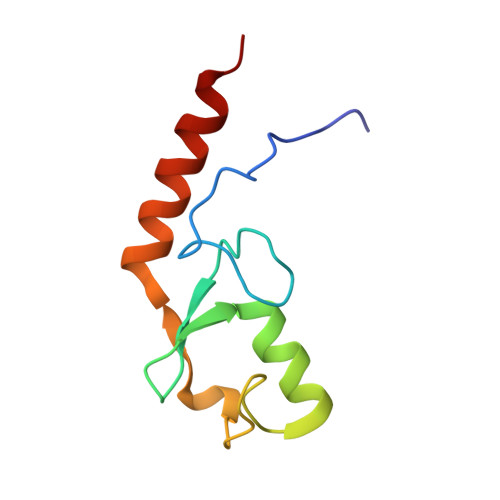
Structural and functional characterization of the monomeric U-box domain from E4B.
Nordquist, K.A., Dimitrova, Y.N., Brzovic, P.S., Ridenour, W.B., Munro, K.A., Soss, S.E., Caprioli, R.M., Klevit, R.E., Chazin, W.J.(2010) Biochemistry 49: 347-355
- PubMed: 20017557 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi901620v
- Primary Citation Related Structures:
2KR4 - PubMed Abstract:
Substantial evidence has accumulated indicating a significant role for oligomerization in the function of E3 ubiquitin ligases. Among the many characterized E3 ligases, the yeast U-box protein Ufd2 and its mammalian homologue E4B appear to be unique in functioning as monomers. An E4B U-box domain construct (E4BU) has been subcloned, overexpressed in Escherichia coli, and purified, which enabled determination of a high-resolution NMR solution structure and detailed biophysical analysis. E4BU is a stable monomeric protein that folds into the same structure observed for other structurally characterized U-box domain homodimers. Multiple sequence alignment combined with comparative structural analysis reveals substitutions in the sequence that inhibit dimerization. The interaction between E4BU and the E2 conjugating enzyme UbcH5c has been mapped using NMR, and these data have been used to generate a structural model for the complex. The E2 binding site is found to be similar to that observed for dimeric U-box and RING domain E3 ligases. Despite the inability to dimerize, E4BU was found to be active in a standard autoubiquitination assay. The structure of E4BU and its ability to function as a monomer are discussed in light of the ubiquitous observation of U-box and RING domain oligomerization.
- Department of Biochemistry, Vanderbilt University, Nashville,Tennessee 37232, USA.
Organizational Affiliation: