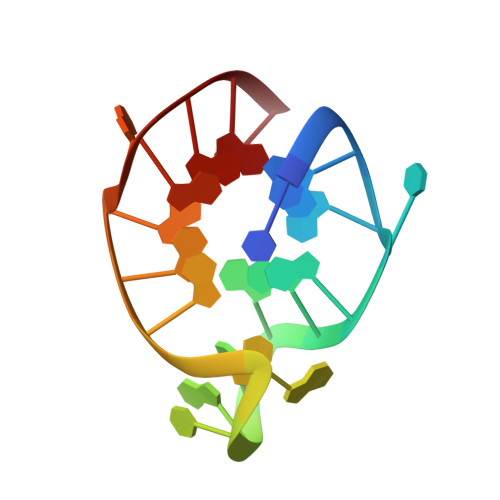
A G-rich sequence within the c-kit oncogene promoter forms a parallel G-quadruplex having asymmetric G-tetrad dynamics
Hsu, S.-T.D., Varnai, P., Bugaut, A., Reszka, A.P., Neidle, S., Balasubramanian, S.(2009) J Am Chem Soc 131: 13399-13409
- PubMed: 19705869 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/ja904007p
- Primary Citation Related Structures:
2KQG, 2KQH - PubMed Abstract:
Guanine-rich DNA sequences with the ability to form quadruplex structures are enriched in the promoter regions of protein-coding genes, particularly those of proto-oncogenes. G-quadruplexes are structurally polymorphic and their folding topologies can depend on the sample conditions. We report here on a structural study using solution state NMR spectroscopy of a second G-quadruplex-forming motif (c-kit2) that has been recently identified in the promoter region of the c-kit oncogene. In the presence of potassium ions, c-kit2 exists as an ensemble of structures that share the same parallel-stranded propeller-type conformations. Subtle differences in structural dynamics have been identified using hydrogen-deuterium exchange experiments by NMR spectroscopy, suggesting the coexistence of at least two structurally similar but dynamically distinct substates, which undergo slow interconversion on the NMR timescale.
- University Chemical Laboratory, University of Cambridge, Lensfield Road, Cambridge, CB2 1EW, United Kingdom.
Organizational Affiliation: