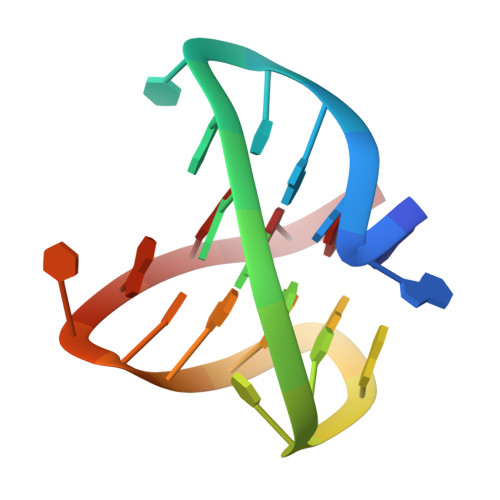
Sequence variant (CTAGGG)n in the human telomere favors a G-quadruplex structure containing a G.C.G.C tetrad
Lim, K.W., Alberti, P., Guedin, A., Lacroix, L., Riou, J.F., Royle, N.J., Mergny, J.L., Phan, A.T.(2009) Nucleic Acids Res 37: 6239-6248
- PubMed: 19692585 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkp630
- Primary Citation Related Structures:
2KM3 - PubMed Abstract:
Short contiguous arrays of variant CTAGGG repeats in the human telomere are unstable in the male germline and somatic cells, suggesting formation of unusual structures by this repeat type. Here, we report on the structure of an intramolecular G-quadruplex formed by DNA sequences containing four human telomeric variant CTAGGG repeats in potassium solution. Our results reveal a new robust antiparallel G-quadruplex fold involving two G-tetrads sandwiched between a G.C base pair and a G.C.G.C tetrad, which could represent a new platform for drug design targeted to human telomeric DNA.
- Division of Physics and Applied Physics, School of Physical and Mathematical Sciences, Nanyang Technological University, Singapore.
Organizational Affiliation: