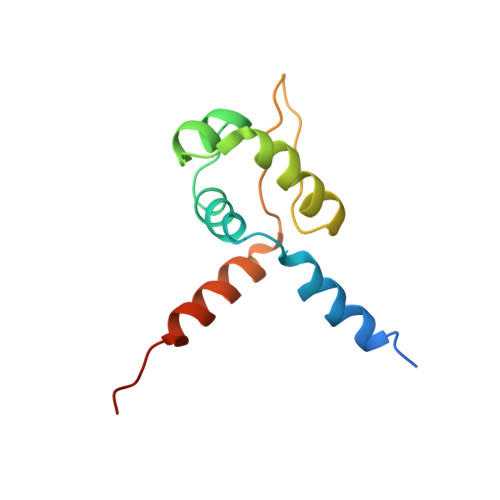
Solution structure of a paradigm ArsR family zinc sensor in the DNA-bound state
Arunkumar, A.I., Campanello, G.C., Giedroc, D.P.(2009) Proc Natl Acad Sci U S A 106: 18177-18182
- PubMed: 19822742 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0905558106
- Primary Citation Related Structures:
2KJB, 2KJC - PubMed Abstract:
Staphylococcus aureus CzrA is a zinc-dependent transcriptional repressor from the ubiquitous ArsR family of metal sensor proteins. Zn(II) binds to a pair of intersubunit C-terminal alpha5-sensing sites, some 15 A distant from the DNA-binding interface, and allosterically inhibits DNA binding. This regulation is characterized by a large allosteric coupling free energy (DeltaGc) of approximately +6 kcal mol(-1), the molecular origin of which is poorly understood. Here, we report the solution quaternary structure of homodimeric CzrA bound to a palindromic 28-bp czr operator, a structure that provides an opportunity to compare the two allosteric "end" states of an ArsR family sensor. Zn(II) binding drives a quaternary structural switch from a "closed" DNA-binding state to a low affinity "open" conformation as a result of a dramatic change in the relative orientations of the winged helical DNA binding domains within the dimer. Zn(II) binding also effectively quenches both rapid and intermediate timescale internal motions of apo-CzrA while stabilizing the native state ensemble. In contrast, DNA binding significantly enhances protein motions in the allosteric sites and reduces the stability of the alpha5 helices as measured by H-D solvent exchange. This study reveals how changes in the global structure and dynamics drive a long-range allosteric response in a large subfamily of bacterial metal sensor proteins, and provides insights on how other structural classes of ArsR sensor proteins may be regulated by metal binding.
- Department of Chemistry, Indiana University, Bloomington, IN 47405-7102, USA.
Organizational Affiliation: