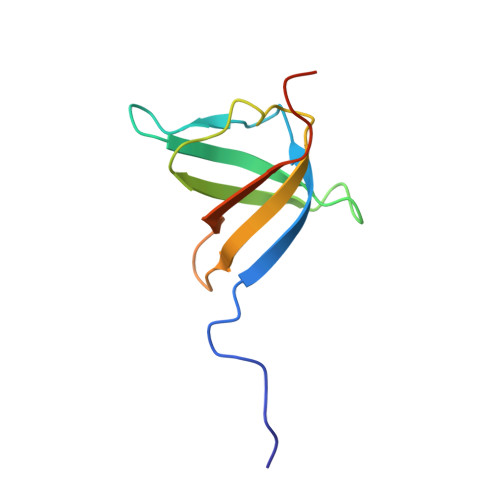
Solution nmr structure of the ob-fold domain of heme chaperone ccme from desulfovibrio vulgaris. northeast structural genomics target dvr115g.
Aramini, J.M., Rossi, P., Lee, H., Lemak, A., Wang, H., Foote, E.L., Jiang, M., Xiao, R., Nair, R., Swapna, G.V.T., Acton, T.B., Rost, B., Everett, J.K., Montelione, G.T.To be published.