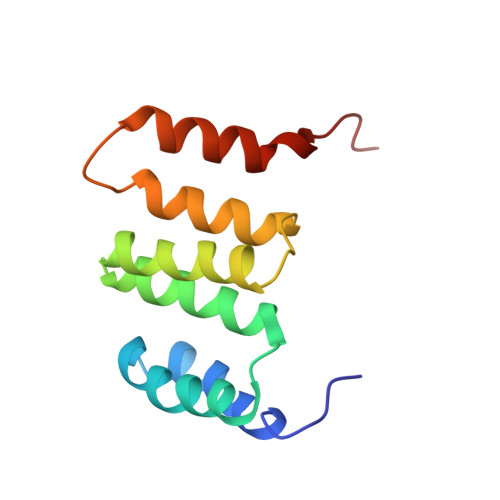
Solution NMR structure of the Northeast Structural Genomics Target MrR121A
Barb, A.W., Lee, H.-W., Wang, X., Lee, D., Jiang, M., Ciccosanti, C., Xiao, R., Nair, R., Everett, J.K., Swapna, G.V.T., Acton, T.B., Rost, B., Montelione, G.T., Prestegard, J.H.To be published.