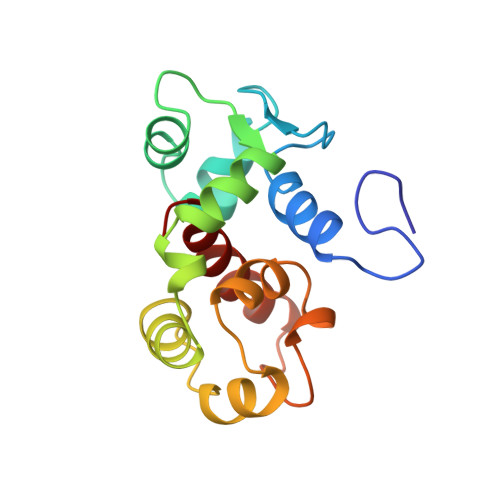
Structural and mechanistic insights into STIM1-mediated initiation of store-operated calcium entry.
Stathopulos, P.B., Zheng, L., Li, G.Y., Plevin, M.J., Ikura, M.(2008) Cell 135: 110-122
- PubMed: 18854159 Search on PubMed
- DOI: https://doi.org/10.1016/j.cell.2008.08.006
- Primary Citation Related Structures:
2K60 - PubMed Abstract:
Stromal interaction molecule-1 (STIM1) activates store-operated Ca2+ entry (SOCE) in response to diminished luminal Ca2+ levels. Here, we present the atomic structure of the Ca2+-sensing region of STIM1 consisting of the EF-hand and sterile alpha motif (SAM) domains (EF-SAM). The canonical EF-hand is paired with a previously unidentified EF-hand. Together, the EF-hand pair mediates mutually indispensable hydrophobic interactions between the EF-hand and SAM domains. Structurally critical mutations in the canonical EF-hand, "hidden" EF-hand, or SAM domain disrupt Ca2+ sensitivity in oligomerization via destabilization of the entire EF-SAM entity. In mammalian cells, EF-SAM destabilization mutations within full-length STIM1 induce punctae formation and activate SOCE independent of luminal Ca2+. We provide atomic resolution insight into the molecular basis for STIM1-mediated SOCE initiation and show that the folded/unfolded state of the Ca2+-sensing region of STIM is crucial to SOCE regulation.
- Division of Signaling Biology, Ontario Cancer Institute and Department of Medical Biophysics, University of Toronto, Toronto Medical Discovery Tower, MaRS Centre, 101 College Street, Toronto, Ontario M5G 1L7, Canada.
Organizational Affiliation: