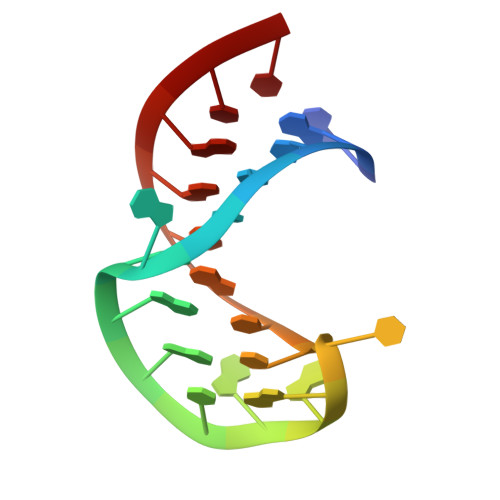
Solution structure of stem-loop alpha of the hepatitis B virus post-transcriptional regulatory element
Schwalbe, M., Ohlenschlager, O., Marchanka, A., Ramachandran, R., Hafner, S., Heise, T., Gorlach, M.(2008) Nucleic Acids Res 36: 1681-1689
- PubMed: 18263618 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkn006
- Primary Citation Related Structures:
2JYM - PubMed Abstract:
Chronic hepatitis B virus (HBV) infections may lead to severe diseases like liver cirrhosis or hepatocellular carcinoma (HCC). The HBV post-transcriptional regulatory element (HPRE) facilitates the nuclear export of unspliced viral mRNAs, contains a splicing regulatory element and resides in the 3'-region of all viral transcripts. The HPRE consists of three sub-elements alpha (nucleotides 1151-1346), beta1 (nucleotides 1347-1457) and beta2 (nucleotides 1458-1582), which confer together full export competence. Here, we present the NMR solution structure (pdb 2JYM) of the stem-loop alpha (SLalpha, nucleotides 1292-1321) located in the sub-element alpha. The SLalpha contains a CAGGC pentaloop highly conserved in hepatoviruses, which essentially adopts a CUNG-like tetraloop conformation. Furthermore, the SLalpha harbours a single bulged G residue flanked by A-helical regions. The structure is highly suggestive of serving two functions in the context of export of unspliced viral RNA: binding sterile alpha motif (SAM-) domain containing proteins and/or preventing the utilization of a 3'-splice site contained within SLalpha.
- Leibniz-Institut für Altersforschung/Fritz-Lipmann-Institut, Beutenbergstr. 11, D-07745 Jena, Germany.
Organizational Affiliation: