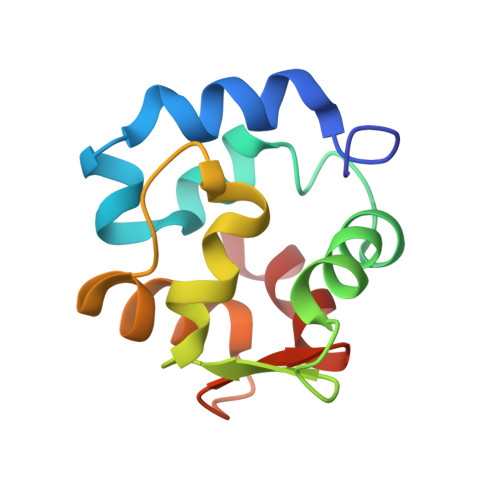
Solution structure of Ca2+-free rat alpha-parvalbumin
Henzl, M.T., Tanner, J.J.(2008) Protein Sci 17: 431-438
- PubMed: 18218708 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.073318308
- Primary Citation Related Structures:
2JWW - PubMed Abstract:
Mammals express two parvalbumins-an alpha isoform and a beta isoform. In rat, the alpha-parvalbumin (alpha-PV) exhibits superior divalent ion affinity. For example, the standard free energies for Ca2+ binding differ by 5.5 kcal/mol in 0.15 M KCl (pH 7.4). High-resolution structures of the Ca2+-bound proteins provide little insight into this disparity, prompting a structural analysis of the apo-proteins. A recent analysis of rat beta-PV suggested that Ca2+ removal provokes substantial conformational changes-reorientation of the C, D, and E helices; reorganization of the hydrophobic core; reduced interdomain contact; and remodeling of the AB domain. The energetic penalty attendant to reversing these changes, it was suggested, could contribute to the attenuated divalent ion-binding signature of that protein. That hypothesis is supported by data presented herein, describing the solution structure and peptide backbone dynamics of Ca2+-free rat alpha-PV. In marked contrast to rat beta-PV, the apo- and Ca2+-loaded forms of the rat alpha isoform are quite similar. Significant structural differences appear to be confined to the loop regions of the molecule. This finding implies that the alpha-PV isoform enjoys elevated divalent ion affinity because the metal ion-binding events do not require major structural rearrangement and the concomitant sacrifice of binding energy.
- Department of Biochemistry, University of Missouri-Columbia, Columbia, Missouri 65211, USA. henzlm@missouri.edu
Organizational Affiliation: