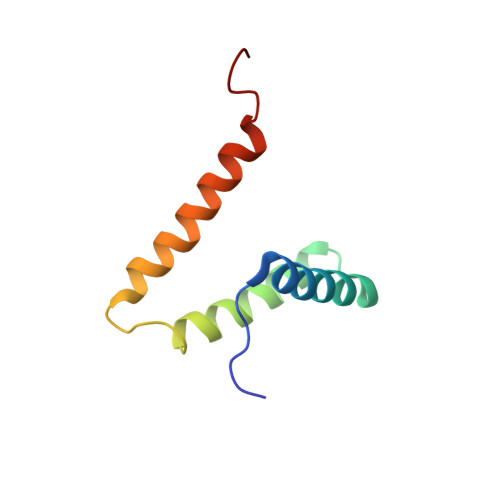
Solution NMR Structure of HI0947 from Haemophilus influenzae.
Ding, K., Ramelot, T.A., Cort, J.R., Wang, D., Nwosu, C., Owens, L., Xiao, R., Liu, J., Baran, M.C., Swapna, G.V.T., Acton, T.B., Rost, B., Montelione, G.T., Kennedy, M.A.To be published.