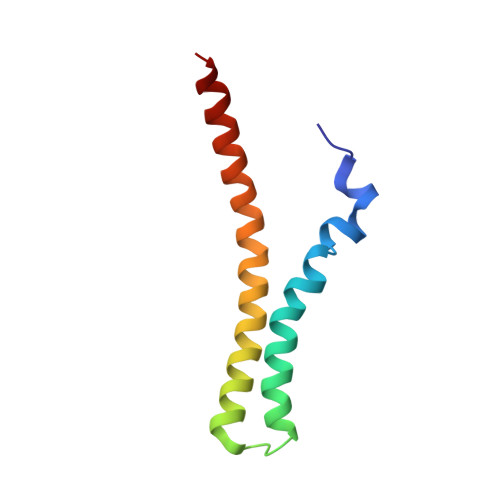
Differences in the Electrostatic Surfaces of the Type III Secretion Needle Proteins PrgI, BsaL, and MxiH
Wang, Y., Ouellette, A.N., Egan, C.W., Rathinavelan, T., Im, W., De Guzman, R.N.(2007) J Mol Biology 371: 1304-1314
- PubMed: 17617421 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2007.06.034
- Primary Citation Related Structures:
2JOW - PubMed Abstract:
Gram-negative bacteria use a needle-like protein assembly, the type III secretion apparatus, to inject virulence factors into target cells to initiate human disease. The needle is formed by the polymerization of approximately 120 copies of a small acidic protein that is conserved among diverse pathogens. We previously reported the structure of the BsaL needle monomer from Burkholderia pseudomallei by nuclear magnetic resonance (NMR) spectroscopy and others have determined the crystal structure of the Shigella flexneri MxiH needle. Here, we report the NMR structure of the PrgI needle protein of Salmonella typhimurium, a human pathogen associated with food poisoning. PrgI, BsaL, and MxiH form similar two helix bundles, however, the electrostatic surfaces of PrgI differ radically from those of BsaL or MxiH. In BsaL and MxiH, a large negative area is on a face formed by the helix alpha1-alpha2 interface. In PrgI, the major negatively charged surface is not on the "face" but instead is on the "side" of the two-helix bundle, and only residues from helix alpha1 contribute to this negative region. Despite being highly acidic proteins, these molecules contain large basic regions, suggesting that electrostatic contacts are important in needle assembly. Our results also suggest that needle-packing interactions may be different among these bacteria and provide the structural basis for why PrgI and MxiH, despite 63% sequence identity, are not interchangeable in S. typhimurium and S. flexneri.
- Department of Molecular Biosciences, The University of Kansas, 1200 Sunnyside Avenue, Lawrence, KA 66045, USA.
Organizational Affiliation: