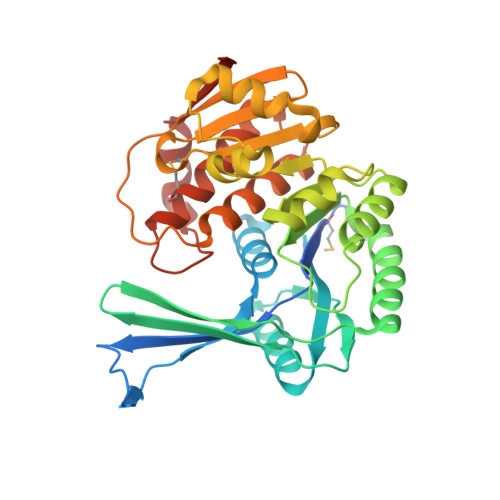
Structures of Staphylococcus Aureus D-Tagatose-6-Phosphate Kinase Implicate Domain Motions in Specificity and Mechanism.
Miallau, L., Hunter, W.N., Mcsweeney, S.M., Leonard, G.A.(2007) J Biological Chem 282: 19948
- PubMed: 17459874 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M701480200
- Primary Citation Related Structures:
2JG1, 2JGV - PubMed Abstract:
High resolution structures of Staphylococcus aureus d-tagatose-6-phosphate kinase (LacC) in two crystal forms are herein reported. The structures define LacC in apoform, in binary complexes with ADP or the co-factor analogue AMP-PNP, and in a ternary complex with AMP-PNP and D-tagatose-6-phosphate. The tertiary structure of the LacC monomer, which is closely related to other members of the pfkB subfamily of carbohydrate kinases, is composed of a large alpha/beta core domain and a smaller, largely beta "lid." Four extended polypeptide segments connect these two domains. Dimerization of LacC occurs via interactions between lid domains, which come together to form a beta-clasp structure. Residues from both subunits contribute to substrate binding. LacC adopts a closed structure required for phosphoryl transfer only when both substrate and co-factor are bound. A reaction mechanism similar to that used by other phosphoryl transferases is proposed, although unusually, when both substrate and co-factor are bound to the enzyme two Mg(2+) ions are observed in the active site. A new motif of amino acid sequence conservation common to the pfkB subfamily of carbohydrate kinases is identified.
- Macromolecular Crystallography Group, European Synchrotron Radiation Facility, 38043 Grenoble, France.
Organizational Affiliation: