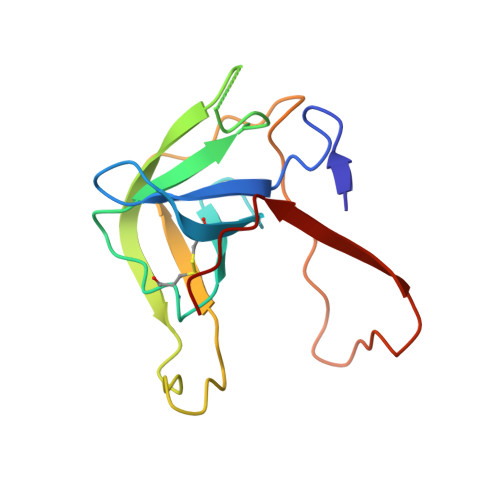
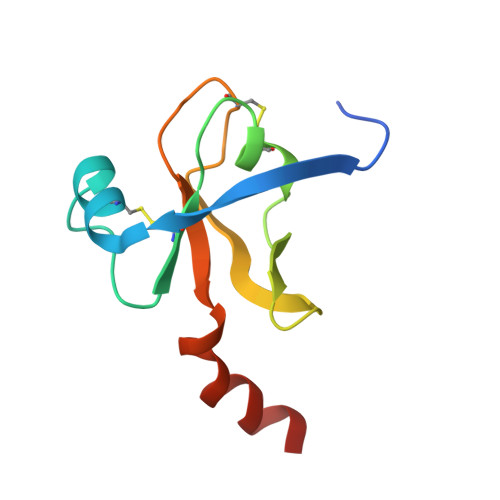
The Crystal Structure of a Trypsin-Like Mutant Chymotrypsin: The Role of Position 226 in the Activity and Specificity of S189D Chymotrypsin.
Jelinek, B., Katona, G., Fodor, K., Venekei, I., Graf, L.(2008) Protein J 27: 79
- PubMed: 17805946 Search on PubMed
- DOI: https://doi.org/10.1007/s10930-007-9110-3
- Primary Citation Related Structures:
2JET - PubMed Abstract:
The crystal structure of the S189D+A226G rat chymotrypsin-B mutant has been determined at 2.2 angstroms resolution. This mutant is the most trypsin-like mutant so far in the line of chymotrypsin-to-trypsin conversions, aiming for a more complete understanding of the structural basis of substrate specificity in pancreatic serine proteases. A226G caused significant rearrangements relative to S189D chymotrypsin, allowing an internal conformation of Asp189 which is close to that in trypsin. Serious distortions remain, however, in the activation domain, including zymogen-like features. The pH-profile of activity suggests that the conformation of the S1-site of the mutant is influenced also by the P1 residue of the substrate.
- Department of Biochemistry, Eötvös Loránd University, Pázmány s. 1/C, Budapest, 1117, Hungary. jelinek@elte.hu
Organizational Affiliation: