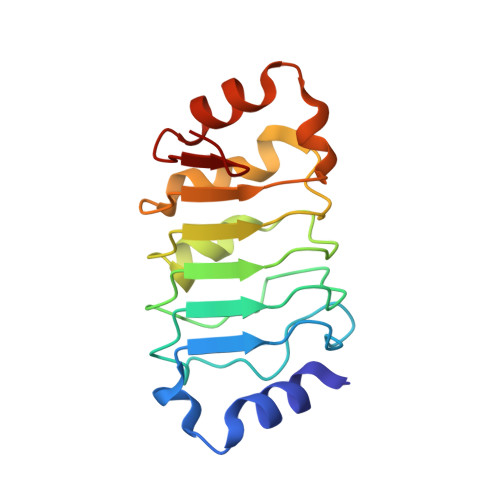
The Crystal Structure of the Tumor Suppressor Protein Pp32 (Anp32A): Structural Insights Into Anp32 Family of Proteins.
Huyton, T., Wolberger, C.(2007) Protein Sci 16: 1308
- PubMed: 17567741 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.072803507
- Primary Citation Related Structures:
2JE0, 2JE1 - PubMed Abstract:
The tumor suppressor protein pp32 is highly overexpressed in many cancers of the breast and prostate, and has also been implicated in the neurodegenerative disease spinocerebellar ataxias type 1 (SCA1). Pp32 is a multifunctional protein that is involved in the regulation of transcription, apoptosis, phosphorylation, and cell cycle progression, the latter through its association with the hyperphosphorylated form of the retinoblastoma tumor suppressor. We have determined the structure of an N-terminal pp32 fragment comprising a capped leucine-rich repeat (LRR) domain, which provides insight into the structural and biochemical properties of the pp32 (Anp32) family of proteins.
- Department of Biophysics and Biophysical Chemistry, Howard Hughes Medical Institute, Johns Hopkins University School of Medicine, Baltimore, MD 21205-2185, USA.
Organizational Affiliation: