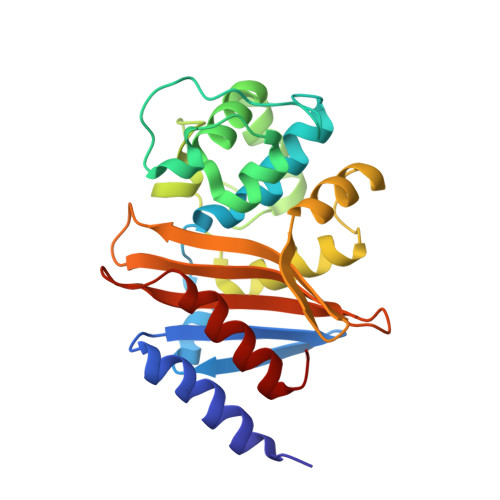
Crystal Structure of the Carbapenemase Oxa-24 Reveals Insights Into the Mechanism of Carbapenem Hydrolysis.
Santillana, E., Beceiro, A., Bou, G., Romero, A.(2007) Proc Natl Acad Sci U S A 104: 5354
- PubMed: 17374723 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0607557104
- Primary Citation Related Structures:
2JC7 - PubMed Abstract:
Combating bacterial resistance to beta-lactams, the most widely used antibiotics, is an emergent and clinically important challenge. OXA-24 is a class D beta-lactamase isolated from a multiresistant epidemic clinical strain of Acinetobacter baumannii. We have investigated how OXA-24 specifically hydrolyzes the last resort carbapenem antibiotic, and we have determined the crystal structure of OXA-24 at a resolution of 2.5 A. The structure shows that the carbapenem's substrate specificity is determined by a hydrophobic barrier that is established through the specific arrangement of the Tyr-112 and Met-223 side chains, which define a tunnel-like entrance to the active site. The importance of these residues was further confirmed by mutagenesis studies. Biochemical and microbiological analyses of specific point mutants selected on the basis of structural criteria significantly reduced the catalytic efficiency (k(cat)/K(m)) against carbapenems, whereas the specificity for oxacillin was noticeably increased. This is the previously unrecognized crystal structure that has been obtained for a class D carbapenemase enzyme. Accordingly, this information may help to improve the development of effective new drugs to combat beta-lactam resistance. More specifically, it may help to overcome carbapenem resistance in A. baumannii, probably one of the most worrying infectious threats in hospitals worldwide.
- Departamento de Estructura y Función de Proteínas, Centro de Investigaciones Biológicas, Consejo Superior de Investigaciones Cientificas, Ramiro de Maeztu 9, E-28040 Madrid, Spain.
Organizational Affiliation: