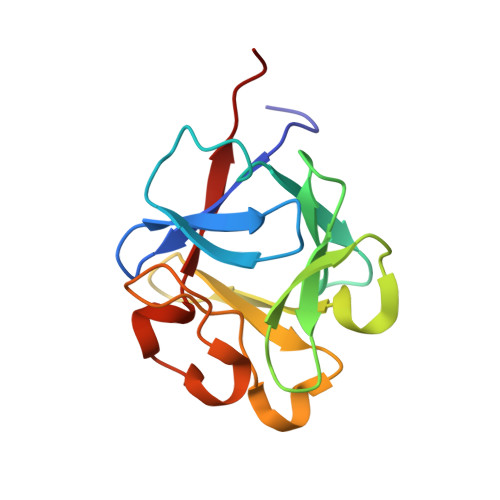
Structure of Rat Acidic Fibroblast Growth Factor at 1.4 A Resolution.
Kulahin, N., Kiselyov, V., Kochoyan, A., Kristensen, O., Kastrup, J.S., Berezin, V., Bock, E., Gajhede, M.(2007) Acta Crystallogr Sect F Struct Biol Cryst Commun 63: 65
- PubMed: 17277441 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309107003144
- Primary Citation Related Structures:
2J3P - PubMed Abstract:
Fibroblast growth factors (FGFs) constitute a family of 22 structurally related heparin-binding polypeptides that are involved in the regulation of cell growth, survival, differentiation and migration. Here, a 1.4 A resolution X-ray structure of rat FGF1 is presented. Two molecules are present in the asymmetric unit of the crystal and they coordinate a total of five sulfate ions. The structures of human, bovine and newt FGF1 have been published previously. Human and rat FGF1 are found to have very similar structures.
- Protein Laboratory, Institute of Molecular Pathology, Panum Institute, Blegdamsvej 3C, DK-2200 Copenhagen, Denmark. kulahin@plab.ku.dk
Organizational Affiliation: