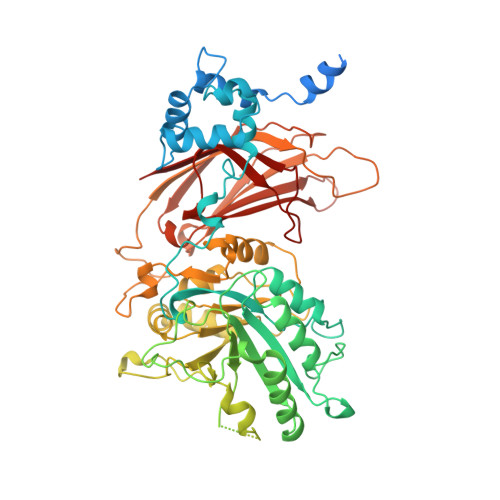
Crystal structure of a mammalian phosphoinositide-specific phospholipase C delta.
Essen, L.O., Perisic, O., Cheung, R., Katan, M., Williams, R.L.(1996) Nature 380: 595-602
- PubMed: 8602259 Search on PubMed
- DOI: https://doi.org/10.1038/380595a0
- Primary Citation Related Structures:
2ISD - PubMed Abstract:
Mammalian phosphoinositide-specific phospholipase C enzymes (PI-PLC) act as signal transducers that generate two second messengers, inositol-1,4,5-trisphosphate and diacylglycerol. The 2.4-A structure of phospholipase C delta 1 reveals a multidomain protein incorporating modules shared by many signalling proteins. The structure suggests a mechanism for membrane attachment and Ca2+-dependent hydrolysis of second-messenger precursors. The regulation and reversible membrane association of PI-PLC may serve as a model for understanding other multidomain enzymes involved in phospholipid signalling.
- Center for Protein Engineering, MRC Centre, Cambridge, UK.
Organizational Affiliation: