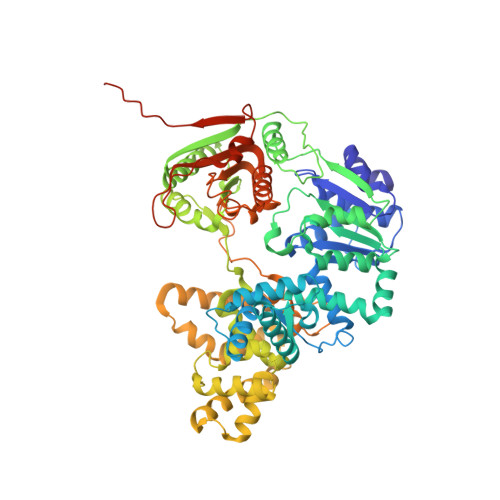
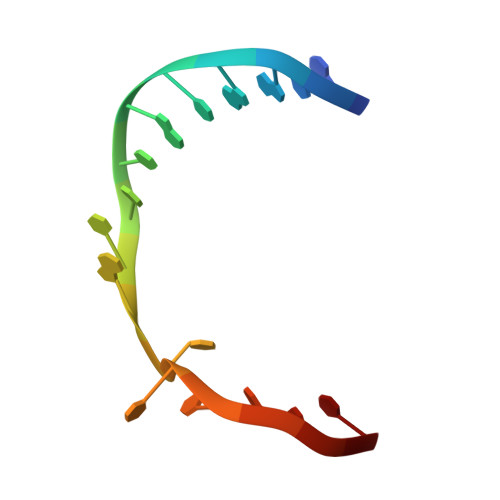
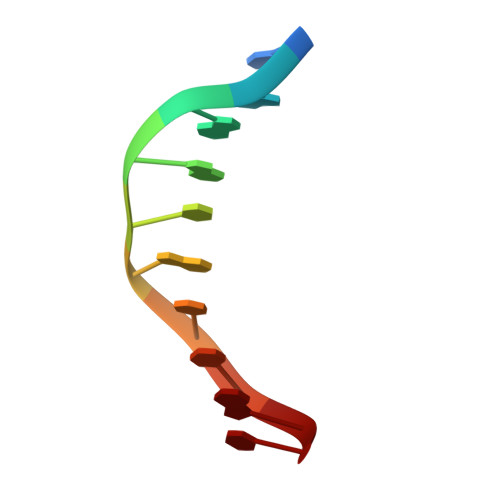
UvrD helicase unwinds DNA one base pair at a time by a two-part power stroke.
Lee, J.Y., Yang, W.(2006) Cell 127: 1349-1360
- PubMed: 17190599 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.cell.2006.10.049
- Primary Citation Related Structures:
2IS1, 2IS2, 2IS4, 2IS6 - PubMed Abstract:
Helicases use the energy derived from nucleoside triphosphate hydrolysis to unwind double helices in essentially every metabolic pathway involving nucleic acids. Earlier crystal structures have suggested that DNA helicases translocate along a single-stranded DNA in an inchworm fashion. We report here a series of crystal structures of the UvrD helicase complexed with DNA and ATP hydrolysis intermediates. These structures reveal that ATP binding alone leads to unwinding of 1 base pair by directional rotation and translation of the DNA duplex, and ADP and Pi release leads to translocation of the developing single strand. Thus DNA unwinding is achieved by a two-part power stroke in a combined wrench-and-inchworm mechanism. The rotational angle and translational distance of DNA define the unwinding step to be 1 base pair per ATP hydrolyzed. Finally, a gateway for ssDNA translocation and an alternative strand-displacement mode may explain the varying step sizes reported previously.
- Laboratory of Molecular Biology, National Institute of Diabetes and Digestive and Kidney Diseases, National Institutes of Health, Bethesda, MD 20892, USA.
Organizational Affiliation: